 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AA97G00029 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Aethionemeae; Aethionema
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 245aa MW: 28467 Da PI: 9.1364 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 53.9 | 4e-17 | 10 | 57 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT++Ed lv+ v+++G ++W+ Ia+ g++Rt+k+c++rw +yl
AA97G00029 10 KGPWTEQEDMVLVNFVHLFGDRRWDFIAKVSGLNRTGKSCRLRWVNYL 57
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 54.3 | 3e-17 | 63 | 108 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T +E+ l +++++++G++ W++Iar+++ gRt++++k++w++++
AA97G00029 63 RGKMTSQEERLVLELHAKWGNR-WSKIARKLP-GRTDNEIKNYWRTHM 108
89********************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 17.36 | 5 | 57 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 5.03E-30 | 8 | 104 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.4E-15 | 9 | 59 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 9.7E-16 | 10 | 57 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 8.8E-21 | 11 | 64 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 6.08E-11 | 12 | 57 | No hit | No description |
| PROSITE profile | PS51294 | 26.962 | 58 | 112 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.9E-16 | 62 | 110 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.2E-15 | 63 | 107 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.0E-23 | 65 | 111 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.35E-11 | 65 | 108 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 245 aa Download sequence Send to blast |
MKIMQEDFRK GPWTEQEDMV LVNFVHLFGD RRWDFIAKVS GLNRTGKSCR LRWVNYLHPG 60 LKRGKMTSQE ERLVLELHAK WGNRWSKIAR KLPGRTDNEI KNYWRTHMRK KAQEKKRPIS 120 RTSSSSNCCS SSMTTATTQE TSPRFEKESF YDTGGSYGKM NQEEGYSMDD IWREIDQSGG 180 NLMKPVRDIY SEQSCYLNCP SMASPAWESS LESIWNMDAY ESKISSFGFD QFPVSFGHSR 240 SSLAD |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1h88_C | 2e-29 | 2 | 112 | 50 | 159 | MYB PROTO-ONCOGENE PROTEIN |
| 1h89_C | 2e-29 | 2 | 112 | 50 | 159 | MYB PROTO-ONCOGENE PROTEIN |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. {ECO:0000305}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00600 | PBM | Transfer from AT5G59780 | Download |
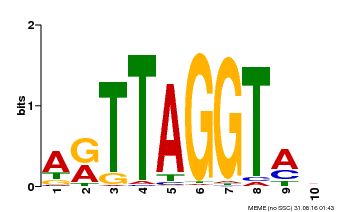
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AA97G00029 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Isoform MYB59-1 is induced by jasmonate (JA), salicylic acid (SA), gibberellic acid (GA), and ethylene. Also induced by cadmium (Cd). {ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:16531467}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010443755.1 | 1e-137 | PREDICTED: transcription factor MYB59 isoform X1 | ||||
| Swissprot | Q4JL84 | 1e-130 | MYB59_ARATH; Transcription factor MYB59 | ||||
| TrEMBL | A0A078JWX0 | 1e-132 | A0A078JWX0_BRANA; BnaAnng32620D protein | ||||
| TrEMBL | A0A398A807 | 1e-132 | A0A398A807_BRACM; Uncharacterized protein | ||||
| TrEMBL | M4DUU6 | 1e-132 | M4DUU6_BRARP; Uncharacterized protein | ||||
| STRING | XP_010443755.1 | 1e-137 | (Camelina sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2502 | 27 | 74 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G59780.3 | 1e-128 | myb domain protein 59 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



