 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AA4G00058 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Aethionemeae; Aethionema
|
||||||||
| Family | C3H | ||||||||
| Protein Properties | Length: 475aa MW: 53412.3 Da PI: 6.0346 | ||||||||
| Description | C3H family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-CCCH | 44.4 | 2.7e-14 | 97 | 119 | 4 | 26 |
---SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 4 elCrffartGtCkyGdrCkFaHg 26
e C+f++rtGtCkyG++CkF+H+
AA4G00058 97 EDCSFYMRTGTCKYGSTCKFNHP 119
89********************9 PP
| |||||||
| 2 | zf-CCCH | 34.1 | 4.7e-11 | 186 | 206 | 5 | 25 |
--SGGGGTS--TTTTT-SS-S CS
zf-CCCH 5 lCrffartGtCkyGdrCkFaH 25
+C+f++rtG CkyGd+C+F+H
AA4G00058 186 ECKFYFRTGGCKYGDSCRFSH 206
6******************** PP
| |||||||
| 3 | zf-CCCH | 38.9 | 1.4e-12 | 229 | 253 | 2 | 26 |
-S---SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 2 ktelCrffartGtCkyGdrCkFaHg 26
++++C+f++r+G Ck+G+ CkF+H+
AA4G00058 229 GEKECPFYMRNGSCKFGADCKFNHP 253
7899********************9 PP
| |||||||
| 4 | zf-CCCH | 20.6 | 7.7e-07 | 376 | 401 | 2 | 27 |
-S---SGGGGTS--TTTTT-SS-SSS CS
zf-CCCH 2 ktelCrffartGtCkyGdrCkFaHgp 27
++++C+++++tG Ck+ +Ck++H++
AA4G00058 376 DQPECSYYLKTGDCKFKYKCKYHHPK 401
689*********************96 PP
| |||||||
| 5 | zf-CCCH | 33.6 | 6.7e-11 | 422 | 446 | 2 | 26 |
-S---SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 2 ktelCrffartGtCkyGdrCkFaHg 26
+++ C++++r+G+Ck+G+ C+F H+
AA4G00058 422 DQSMCSHYSRYGICKFGPACRFDHS 446
5678********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF90229 | 1.04E-6 | 91 | 121 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 4.6E-7 | 93 | 120 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 14.558 | 93 | 121 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 1.2E-11 | 97 | 119 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 1.3E-5 | 99 | 118 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 7.59E-7 | 181 | 209 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 15.469 | 181 | 209 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 8.7E-6 | 181 | 208 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 1.2E-8 | 186 | 206 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 2.5E-6 | 187 | 208 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 16.042 | 227 | 255 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 1.1E-6 | 227 | 254 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 4.3E-10 | 229 | 253 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 7.59E-6 | 229 | 254 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 2.6E-4 | 233 | 252 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 13.028 | 374 | 402 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 0.053 | 374 | 401 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 2.8E-4 | 376 | 401 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 8.9E-5 | 378 | 404 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 14.821 | 420 | 448 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 0.013 | 420 | 447 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 1.57E-5 | 421 | 448 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 1.1E-4 | 424 | 446 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 2.1E-7 | 426 | 446 | IPR000571 | Zinc finger, CCCH-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 475 aa Download sequence Send to blast |
MEKSVEISDP NSSRPDPEST SRSISDEAMI DRGSSDVNHD AEELSDQLKN VGLEETSFRD 60 LGGDDEKERA REEEVGDESE SMMEKRVAVY PLRPEAEDCS FYMRTGTCKY GSTCKFNHPV 120 KRRLQVISSF TNICMFYYFN FPSYDNDFES FWLSIWRKFD LTISGLIGRD RVREREEDVE 180 NPKLMECKFY FRTGGCKYGD SCRFSHTKQK NSLTSGPELN FLGLPIRPGE KECPFYMRNG 240 SCKFGADCKF NHPDPTTIGG TDSPLLRGNG GSVGSFSPKA ASQASSTSWS SSRHMNGTGT 300 GTGTAPFIPV MFSQNRGVSP QTPEWNGYQA ASMYPPERSV LPPSTYMMNN SSAETSLISQ 360 QHQQQIPAEE FPERPDQPEC SYYLKTGDCK FKYKCKYHHP KNRLPKQAPS ALNDKGLPLR 420 PDQSMCSHYS RYGICKFGPA CRFDHSIPPP CLPSSSQTME APQVGNNGNE SDNWN |
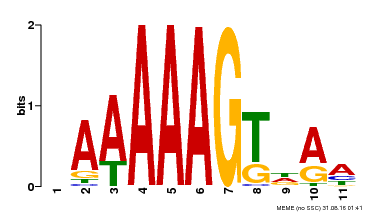
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00582 | DAP | Transfer from AT5G63260 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AA4G00058 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006394295.1 | 0.0 | zinc finger CCCH domain-containing protein 67 isoform X1 | ||||
| Swissprot | Q5RJC5 | 0.0 | C3H67_ARATH; Zinc finger CCCH domain-containing protein 67 | ||||
| TrEMBL | V4MJP0 | 0.0 | V4MJP0_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006394295.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4176 | 27 | 58 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G63260.2 | 0.0 | C3H family protein | ||||



