 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AA21G00382 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Aethionemeae; Aethionema
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 551aa MW: 62117.6 Da PI: 5.0804 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 46.1 | 8.8e-15 | 389 | 435 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr+++P+ + K++Ka+ Le A+ YIk+Lq
AA21G00382 389 NHVEAERQRREKLNQRFYALRSVVPNiS------KMDKASLLEDAIAYIKELQ 435
799***********************66......******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 8.3E-52 | 49 | 237 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| SuperFamily | SSF47459 | 5.76E-19 | 384 | 445 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 18.019 | 385 | 434 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 2.35E-16 | 388 | 439 | No hit | No description |
| Pfam | PF00010 | 4.0E-12 | 389 | 435 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 5.4E-18 | 389 | 445 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 9.3E-18 | 391 | 440 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009611 | Biological Process | response to wounding | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 551 aa Download sequence Send to blast |
MNNISDVGWD DEDKSLVTTV LGHLAYDFLR ANSGSNQNLF LVMGTDDSLH TKLSSLVDWP 60 NSENFSWNYA IFWQQTMSRS GQQVLGWGDG CCREPNEEED SKVIRVYNLG LDVETWEGMR 120 KRVLQKLHRL FGGSDEDNYA KSLEKVTATE MFFLSSMYFF FPHGEGGPGR CYASGKHIWI 180 SDAMTSDSDY CFRSFMAKSA GIRTIVLVPT DAGVLELGSV WSLPENFELV KSVQTLFMRR 240 PRHMNDTVIN GSKTDGIHKL FGQELNNNSE QVVNNGYPKK LNVSQAWEGF QGDSRNGFRG 300 FTCQEMDGKV QELVNGVLDE TRTSSYQSLR LSNQTQIEFT CSVAGSTNTQ LEKSDSCTEK 360 KAIIECNLDE KRPRKRGRKP ANGREEPLNH VEAERQRREK LNQRFYALRS VVPNISKMDK 420 ASLLEDAIAY IKELQEKVKI LESESLSESN QTRTVESPEI DIQAMNDEVV VRVISPLETH 480 PASRVIQAMR NSEVDLMESK LSLAEDTMFH TFVIKNGGDP LTKEKLFAAV YPETSSVQPL 540 PSSSSQVSGD F |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 6e-29 | 383 | 455 | 2 | 74 | Transcription factor MYC2 |
| 5gnj_B | 6e-29 | 383 | 455 | 2 | 74 | Transcription factor MYC2 |
| 5gnj_E | 6e-29 | 383 | 455 | 2 | 74 | Transcription factor MYC2 |
| 5gnj_F | 6e-29 | 383 | 455 | 2 | 74 | Transcription factor MYC2 |
| 5gnj_G | 6e-29 | 383 | 455 | 2 | 74 | Transcription factor MYC2 |
| 5gnj_I | 6e-29 | 383 | 455 | 2 | 74 | Transcription factor MYC2 |
| 5gnj_M | 6e-29 | 383 | 455 | 2 | 74 | Transcription factor MYC2 |
| 5gnj_N | 6e-29 | 383 | 455 | 2 | 74 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 370 | 378 | KRPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator. Regulates positively abscisic acid (ABA) response. Confers drought tolerance and sensitivity to ABA. {ECO:0000269|PubMed:17828375}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
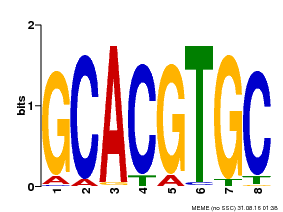
| MP00101 | PBM | Transfer from AT2G46510 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AA21G00382 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Transiently by ABA and polyethylene glycol (PEG). {ECO:0000269|PubMed:17828375}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020884075.1 | 0.0 | transcription factor ABA-INDUCIBLE bHLH-TYPE | ||||
| Swissprot | Q9ZPY8 | 0.0 | AIB_ARATH; Transcription factor ABA-INDUCIBLE bHLH-TYPE | ||||
| TrEMBL | D7LEK3 | 0.0 | D7LEK3_ARALL; Basic helix-loop-helix family protein | ||||
| STRING | fgenesh2_kg.4__2868__AT2G46510.1 | 0.0 | (Arabidopsis lyrata) | ||||
| STRING | Bostr.25993s0115.1.p | 0.0 | (Boechera stricta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3758 | 27 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46510.1 | 0.0 | ABA-inducible BHLH-type transcription factor | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



