 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KFK36045.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Arabideae; Arabis
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 323aa MW: 35447.9 Da PI: 9.5082 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 25.4 | 3.2e-08 | 7 | 57 | 3 | 48 |
SS-HHHHHHHHHHHHHTTTT......-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGgg......tWktIartmgkgRtlkqcksrwqkyl 48
W+ eE++ + +a++++ +W + a++++ +++ +++k ++q +l
KFK36045.1 7 TWSREEEKSFENAIAMHCVQeeitedQWMKMASMVP-TKSIEEVKKHYQLLL 57
7****************999****************.************875 PP
| |||||||
| 2 | Myb_DNA-binding | 38.4 | 2.9e-12 | 141 | 185 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ l++ + ++G+g+W++I+r + Rt+ q+ s+ qky
KFK36045.1 141 PWTEEEHRLFLLGLDKFGKGDWRSISRNFVISRTPTQVASHAQKY 185
7*****************************99************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 8.213 | 3 | 61 | IPR017884 | SANT domain |
| SMART | SM00717 | 0.0095 | 4 | 59 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 7.3E-7 | 7 | 56 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.96E-6 | 7 | 57 | No hit | No description |
| SuperFamily | SSF46689 | 2.55E-8 | 8 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 17.414 | 134 | 190 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.83E-16 | 136 | 191 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.5E-16 | 138 | 188 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 3.0E-10 | 138 | 188 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.6E-11 | 139 | 183 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 1.1E-10 | 141 | 185 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.47E-9 | 141 | 185 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 323 aa Download sequence Send to blast |
MESVVMTWSR EEEKSFENAI AMHCVQEEIT EDQWMKMASM VPTKSIEEVK KHYQLLLEDV 60 KAIENGQVPL PRYHHHHHKK LDDTTTSQVN NKDSPSSGGS GSSEKKPNHG GSGISISNGG 120 GGGGGRNGSS RAEQERRKGI PWTEEEHRLF LLGLDKFGKG DWRSISRNFV ISRTPTQVAS 180 HAQKYYIRLT SMNRDRRRSS IHDITTVNNQ AHAVTGQGGQ QQQQQQVVKH RPTISQSQPQ 240 PQPLPHPHQG AAMGGLGMYG GAPMGQPIIV PPDHMGSAVG TPVMLPPPMG THHHHLGVAA 300 PYPVPAYPVP SLPLQHPAPS TMH |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 4e-13 | 8 | 73 | 11 | 73 | RADIALIS |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00191 | DAP | Transfer from AT1G49010 | Download |
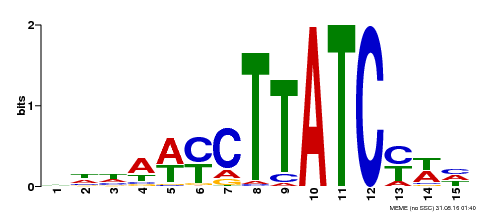
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KFK36045.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC016041 | 9e-62 | AC016041.5 Genomic sequence for Arabidopsis thaliana BAC F27J15 from chromosome I, complete sequence. | |||
| GenBank | AY086906 | 9e-62 | AY086906.1 Arabidopsis thaliana clone 29302 mRNA, complete sequence. | |||
| GenBank | AY519528 | 9e-62 | AY519528.1 Arabidopsis thaliana MYB transcription factor (At1g49010) mRNA, complete cds. | |||
| GenBank | BT024862 | 9e-62 | BT024862.1 Arabidopsis thaliana At1g49010 gene, complete cds. | |||
| GenBank | CP002684 | 9e-62 | CP002684.1 Arabidopsis thaliana chromosome 1 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009147920.1 | 1e-154 | PREDICTED: transcription factor DIVARICATA-like | ||||
| Refseq | XP_018489531.1 | 1e-154 | PREDICTED: transcription factor DIVARICATA-like | ||||
| Refseq | XP_018489584.1 | 1e-154 | PREDICTED: transcription factor DIVARICATA-like | ||||
| TrEMBL | A0A087H1P3 | 0.0 | A0A087H1P3_ARAAL; Uncharacterized protein | ||||
| STRING | A0A087H1P3 | 0.0 | (Arabis alpina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM10304 | 25 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G49010.1 | 1e-107 | MYB family protein | ||||



