 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KFK34139.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Arabideae; Arabis
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1352aa MW: 152071 Da PI: 8.5922 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.5 | 0.00044 | 1235 | 1260 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
++C C++sF++ +L +H r+ +
KFK34139.1 1235 FQCDleGCTMSFNSEKELAVHKRNiC 1260
89********************9877 PP
| |||||||
| 2 | zf-C2H2 | 11.8 | 0.00075 | 1260 | 1282 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L +H r+H
KFK34139.1 1260 CPvkGCGKNFFSHKYLLQHRRVH 1282
9999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 12 | 0.00065 | 1318 | 1344 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
KFK34139.1 1318 YVCAepGCGQTFRFVSDFSRHKRKtgH 1344
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 1.0E-15 | 19 | 60 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 13.786 | 20 | 61 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 7.7E-14 | 21 | 54 | IPR003349 | JmjN domain |
| SMART | SM00558 | 2.2E-51 | 200 | 369 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 33.823 | 203 | 369 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 4.39E-27 | 218 | 387 | No hit | No description |
| Pfam | PF02373 | 1.7E-37 | 234 | 352 | IPR003347 | JmjC domain |
| SMART | SM00355 | 12 | 1235 | 1257 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.902 | 1258 | 1287 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.02 | 1258 | 1282 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1260 | 1282 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 9.0E-6 | 1260 | 1286 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 6.87E-10 | 1274 | 1316 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 9.6E-9 | 1287 | 1311 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0015 | 1288 | 1312 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.928 | 1288 | 1317 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1290 | 1312 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 3.04E-8 | 1306 | 1340 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.8E-9 | 1312 | 1341 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.62 | 1318 | 1344 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.032 | 1318 | 1349 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1320 | 1344 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1352 aa Download sequence Send to blast |
MAVSEQSQDV FPWLKSLPVA PEFRPTLAEF QDPIAYIFKI EEEASRYGIC KILPPVPPPS 60 KKTAISNLNR SLAARAAARV RDGGGSAFDY DGGPTFATRQ QQIGFCPRKQ RPVQRPVWQS 120 GEHYSFGEFE YKAKTFEKNY LKKCSKKSQL SALEMETLYW KATVDKPFSV EYANDMPGSA 180 FIPLSLAAAR RREYGGDGGT VGETAWNMRA MARAEGSLLK FMKEEIPGVT SPMIYIAMMF 240 SWFAWHVEDH DLHSLNYLHM GASKTWYGVP KDAAVAFEEV VRVHGYGGEL NPLVTFSTLG 300 EKTTVMSPEV FVKAGIPCCR LVQNPGDFVV TFPRAYHSGF SHGFNCGEAS NIATPEWLRM 360 AKDAAIRRAA ISYRPIVSHL QLLYDFALAL GSRVPTSIHT KPRSSRLKDK KRSEGEKLTK 420 ELFVQNIIHN NELLHSLGKG SPIALLPQSS SDKSVCSDQR KSLHLYXXXX MVSLSNGVKD 480 TVSVKEKFTS LCERNRNHLE KETQGTLTDA ERRKNVRLSD QRLFSCVTCG ILSFDCVAIV 540 QPKEAAARYL MSADCSFFND WTVASGSANL GQVARSLHPQ SMEKHDADYF YDVPIEIMDQ 600 RTSSTSLTKA QKDNDALGLL ASAYGDSSDS EEEDHKGIDI PISEGAFKAS SVDTDGKEEA 660 RDGRSSDFNS QRLSCGKGKE VEVSHATSTC STLSCTSKQN KLSKGDNTSL VEIALPFVPR 720 SDDDSSRLHV FCLEHAAEVE EQLRPIGGIH IMLLCHPEYP RIEAEAKIVA EELGVNQEWN 780 DTEFRNVTQE DEETIQAALD NVGTKAGNSD WAVKLGINLS YSAILSRSPL YSKQMPYNSI 840 IYNAFGRSSP ATSSPSKLEI SGKRSSRQRK YVVGKWCGKV WMSHQVHPFL LQQDLEGEES 900 DRSHHLRVAL DEDVTGKRLF PGNNSRDATT MFGRKYSRKR KMRAKAVPRK KLTSFKREDG 960 VSDDTSEDHS YKQQWRASDN EEESYFEIGN TGSGDSSNHM SDHQQLKGSR RHKGLKEFES 1020 GDEASDRSLG EEYAVRDYAV SESSMENSFE LYREKQSRYD HEDDDLYRYP RGIPRNKRAK 1080 VFRNPVSYDS EENGAYQQRR RVSTSKQARR MGGEYDSAEN SLEEQNFSVT GKRQTRSTAK 1140 RKAKIEAVQS PSDTEGCGLQ GFASGKKNRE LDSYMEGPST RLRVRNLKPS RGSKVTKPKK 1200 TGRKGISIAT FSRAASEEEL EEDEGENEEE ERTAFQCDLE GCTMSFNSEK ELAVHKRNIC 1260 PVKGCGKNFF SHKYLLQHRR VHSDDRPLKC PWKGCKMTFK WAWSRTEHIR VHTGARPYVC 1320 AEPGCGQTFR FVSDFSRHKR KTGHSVKKTK KR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6a57_A | 3e-74 | 1234 | 1352 | 22 | 140 | Lysine-specific demethylase REF6 |
| 6a58_A | 3e-74 | 1234 | 1352 | 22 | 140 | Lysine-specific demethylase REF6 |
| 6a59_A | 3e-74 | 1234 | 1352 | 22 | 140 | Lysine-specific demethylase REF6 |
| 6ip0_A | 5e-73 | 9 | 465 | 4 | 421 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 5e-73 | 9 | 465 | 4 | 421 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 936 | 941 | SRKRKM |
| 2 | 938 | 950 | KRKMRAKAVPRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
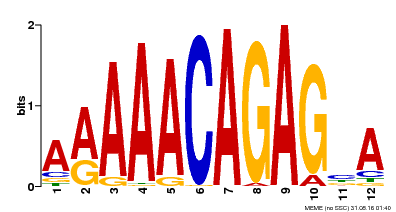
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KFK34139.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY664499 | 0.0 | AY664499.1 Arabidopsis thaliana relative of early flowering 6 (REF6) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006404261.1 | 0.0 | lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A087GW87 | 0.0 | A0A087GW87_ARAAL; Uncharacterized protein | ||||
| STRING | A0A087GW86 | 0.0 | (Arabis alpina) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



