 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp5g01460 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 230aa MW: 27051.2 Da PI: 4.8214 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 56.8 | 3.7e-18 | 32 | 83 | 5 | 56 |
SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 5 ttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+f++eq++ Le Fe++ + +++ +LA++lgL+ rqV +WFqN+Ra++k
Tp5g01460 32 KRFSEEQIKSLEVIFESETRLEPRKKVQLARELGLQPRQVAIWFQNKRARWK 83
68*************************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 118.7 | 3.2e-38 | 30 | 121 | 2 | 93 |
HD-ZIP_I/II 2 kkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
+++r+s+eq+k+LE Fe+e++Lep++Kv+lareLglqprqva+WFqn+RAR+k+kqlEk+y+ L+++y++l+++ e ++ke+++L +el++
Tp5g01460 30 NQKRFSEEQIKSLEVIFESETRLEPRKKVQLARELGLQPRQVAIWFQNKRARWKSKQLEKEYNILRANYNNLASQFEIMKKEKQALVSELQR 121
689*************************************************************************************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 7.52E-18 | 10 | 87 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 7.2E-15 | 22 | 89 | IPR001356 | Homeobox domain |
| PROSITE profile | PS50071 | 17.103 | 25 | 85 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 2.56E-13 | 31 | 86 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-18 | 31 | 92 | IPR009057 | Homeodomain-like |
| Pfam | PF00046 | 1.6E-15 | 32 | 83 | IPR001356 | Homeobox domain |
| PRINTS | PR00031 | 6.5E-6 | 56 | 65 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 60 | 83 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 6.5E-6 | 65 | 81 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 1.5E-15 | 85 | 127 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009615 | Biological Process | response to virus | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 230 aa Download sequence Send to blast |
MEEGDFFNCC FSEINSGMTM NKKKMRKSNN QKRFSEEQIK SLEVIFESET RLEPRKKVQL 60 ARELGLQPRQ VAIWFQNKRA RWKSKQLEKE YNILRANYNN LASQFEIMKK EKQALVSELQ 120 RLNEEMQKSK EERGDECCDK QRVALSSTTR SGSDNGKCEP EVRPNQEIVL CDDVKTEYFG 180 FEEESNHELM NIVEQADDSC LTSSDNWGGF NSECLLDQSS SNYPWWDFWS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription activator that may act as growth regulators in response to water deficit. {ECO:0000269|PubMed:11374882, ECO:0000269|PubMed:15604708}. | |||||
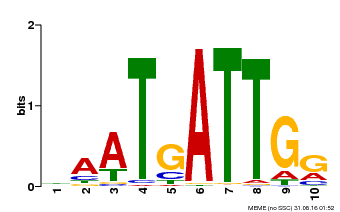
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00013 | PBM | Transfer from AT3G61890 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp5g01460 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By water deficit, by abscisic acid (ABA), by cold and salt stress. {ECO:0000269|PubMed:15369784, ECO:0000269|PubMed:16055682, ECO:0000269|PubMed:9617808}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KT000602 | 0.0 | KT000602.1 Brassica napus note R1 line homeobox-leucine zipper protein (ATHB-12) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009138744.1 | 1e-142 | PREDICTED: homeobox-leucine zipper protein ATHB-12-like | ||||
| Refseq | XP_013631796.1 | 1e-142 | PREDICTED: homeobox-leucine zipper protein ATHB-12-like | ||||
| Refseq | XP_022556559.1 | 1e-142 | homeobox-leucine zipper protein ATHB-12-like | ||||
| Refseq | XP_022556560.1 | 1e-142 | homeobox-leucine zipper protein ATHB-12-like | ||||
| Swissprot | Q9M276 | 1e-116 | ATB12_ARATH; Homeobox-leucine zipper protein ATHB-12 | ||||
| TrEMBL | A0A397ZGR3 | 1e-142 | A0A397ZGR3_BRACM; Uncharacterized protein | ||||
| STRING | Bra014417.1-P | 1e-141 | (Brassica rapa) | ||||
| STRING | Bo4g100070.1 | 1e-141 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2498 | 28 | 74 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G61890.1 | 1e-119 | homeobox 12 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



