 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OBART07G20570.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 219aa MW: 22230.1 Da PI: 8.5917 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 10.9 | 0.0014 | 135 | 157 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C++C ++F++ L H r+H
OBART07G20570.1 135 HRCSICLRTFPSGQALGGHKRLH 157
79*****************9998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50157 | 9.307 | 84 | 111 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.63 | 84 | 106 | IPR015880 | Zinc finger, C2H2-like |
| Pfam | PF13912 | 3.7E-12 | 84 | 108 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.54E-7 | 84 | 157 | No hit | No description |
| PROSITE pattern | PS00028 | 0 | 86 | 106 | IPR007087 | Zinc finger, C2H2 |
| Pfam | PF13912 | 1.2E-12 | 134 | 159 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.27 | 135 | 157 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.162 | 135 | 157 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 137 | 157 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 219 aa Download sequence Send to blast |
MALDGKPPVP PPSTPPMDSW ACGGRRSKRR GGGGGISGSS GSSGGGESEE EYLAACLLML 60 ARGVRDEAEV VGVAAAAKPS QHGYGCSVCG KVYGSYQALG GHKTSHRKPP SPAAEPAAGE 120 EPSSGGMITS EAKVHRCSIC LRTFPSGQAL GGHKRLHYEG GAVGDAVKEK NSLKTKAAAV 180 ATAVLKDFDL NLPAAATTPG DEAESSPPEA KRARLLLLV |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00399 | DAP | Transfer from AT3G49930 | Download |
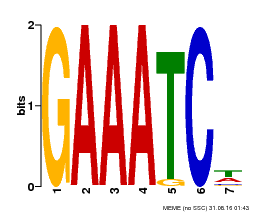
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP005149 | 0.0 | AP005149.4 Oryza sativa Japonica Group genomic DNA, chromosome 7, BAC clone:OSJNBb0005G07. | |||
| GenBank | AP014963 | 0.0 | AP014963.1 Oryza sativa Japonica Group DNA, chromosome 7, cultivar: Nipponbare, complete sequence. | |||
| GenBank | CP012615 | 0.0 | CP012615.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 7 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015647209.1 | 9e-98 | zinc finger protein 1 | ||||
| TrEMBL | A0A0D3GSZ7 | 1e-154 | A0A0D3GSZ7_9ORYZ; Uncharacterized protein | ||||
| STRING | OBART07G20570.1 | 1e-155 | (Oryza barthii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP10028 | 23 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G49930.1 | 6e-25 | C2H2 family protein | ||||



