 |
Plant Transcription
Factor Database
|
Transcription Factor Information
|
Basic
Information? help
Back to Top |
| TF ID |
Cagra.0876s0043.1.p |
| Organism |
|
| Taxonomic ID |
|
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Capsella
|
| Family |
MYB_related |
| Protein Properties |
Length: 210aa MW: 23199.7 Da PI: 5.3331 |
| Description |
MYB_related family protein |
| Gene Model |
| Gene Model ID |
Type |
Source |
Coding Sequence |
| Cagra.0876s0043.1.p | genome | JGI | View CDS |
|
| Signature Domain? help Back to Top |
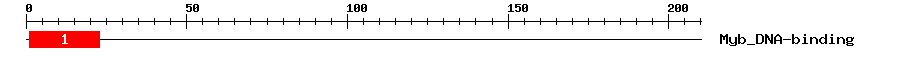 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | Myb_DNA-binding | 29.9 | 1.3e-09 | 1 | 23 | 24 | 47 |
HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 24 WktIartmgkgRtlkqcksrwqky 47
W++Iar+++ gRt++++k++w+++
Cagra.0876s0043.1.p 1 WSAIARRLP-GRTDNEIKNHWNTH 23
*********.************97 PP
|
| Sequence ? help Back to Top |
| Protein Sequence Length: 210 aa
Download sequence Send
to blast |
WSAIARRLPG RTDNEIKNHW NTHIKKRLIK KGIDPVTHKS LLDNLEEKST VHQEKDDQKS 60
KDNKSSRSSA RFLNKVANRF GQRVSQSVLS DIIGSGGSLT SYATPIISSS ECERSTSSSS 120
TPTSSDLLTN QSINVDATTP LSSSTCSDSS DLFLYDDIFG DIEDTTRLIS SRFLSNVVSH 180
DGEEFLMLDG SCLENTSFMR ELTMILQED*
|
| Functional Description ? help
Back to Top |
| Source |
Description |
| UniProt | Transcription factor involved in glucosinolates biosynthesis. {ECO:0000269|PubMed:23580754, ECO:0000269|PubMed:23943862}. |
| Annotation --
Nucleotide ? help
Back to Top |
| Source |
Hit ID |
E-value |
Description |
| GenBank | KJ670120 | 1e-32 | KJ670120.1 Brassica napus MYB transcription factor 122 (MYB122.1) mRNA, complete cds. |
| Publications
? help Back to Top |
- Duarte JM, et al.
Expression pattern shifts following duplication indicative of subfunctionalization and neofunctionalization in regulatory genes of Arabidopsis.
Mol. Biol. Evol., 2006. 23(2): p. 469-78
[PMID:16280546] - Guo R, et al.
Jasmonic acid and glucose synergistically modulate the accumulation of glucosinolates in Arabidopsis thaliana.
J. Exp. Bot., 2013. 64(18): p. 5707-19
[PMID:24151308] - Ding Y, et al.
Four distinct types of dehydration stress memory genes in Arabidopsis thaliana.
BMC Plant Biol., 2013. 13: p. 229
[PMID:24377444] - Frerigmann H,Gigolashvili T
MYB34, MYB51, and MYB122 distinctly regulate indolic glucosinolate biosynthesis in Arabidopsis thaliana.
Mol Plant, 2014. 7(5): p. 814-28
[PMID:24431192] - Frerigmann H,Gigolashvili T
Update on the role of R2R3-MYBs in the regulation of glucosinolates upon sulfur deficiency.
Front Plant Sci, 2014. 5: p. 626
[PMID:25426131] - Peskan-Berghöfer T, et al.
Sustained exposure to abscisic acid enhances the colonization potential of the mutualist fungus Piriformospora indica on Arabidopsis thaliana roots.
New Phytol., 2015. 208(3): p. 873-86
[PMID:26075497] - Frerigmann H,Glawischnig E,Gigolashvili T
The role of MYB34, MYB51 and MYB122 in the regulation of camalexin biosynthesis in Arabidopsis thaliana.
Front Plant Sci, 2015. 6: p. 654
[PMID:26379682] - Frerigmann H, et al.
Regulation of Pathogen-Triggered Tryptophan Metabolism in Arabidopsis thaliana by MYB Transcription Factors and Indole Glucosinolate Conversion Products.
Mol Plant, 2016. 9(5): p. 682-695
[PMID:26802248] - Stahl E, et al.
Regulatory and Functional Aspects of Indolic Metabolism in Plant Systemic Acquired Resistance.
Mol Plant, 2016. 9(5): p. 662-681
[PMID:26802249] - Bulgakov VP,Veremeichik GN,Grigorchuk VP,Rybin VG,Shkryl YN
The rolB gene activates secondary metabolism in Arabidopsis calli via selective activation of genes encoding MYB and bHLH transcription factors.
Plant Physiol. Biochem., 2016. 102: p. 70-9
[PMID:26913794] - Xu J, et al.
Pathogen-Responsive MPK3 and MPK6 Reprogram the Biosynthesis of Indole Glucosinolates and Their Derivatives in Arabidopsis Immunity.
Plant Cell, 2016. 28(5): p. 1144-62
[PMID:27081184] - Miao H, et al.
Glucose enhances indolic glucosinolate biosynthesis without reducing primary sulfur assimilation.
Sci Rep, 2016. 6: p. 31854
[PMID:27549907]
|




