 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 441667 | ||||||||
| Common Name | SELMODRAFT_441667 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Lycopodiidae; Selaginellales; Selaginellaceae; Selaginella
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 983aa MW: 109753 Da PI: 6.6126 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 175.3 | 8.3e-55 | 24 | 138 | 4 | 117 |
CG-1 4 e.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcywlLeee 105
e ++rwl++ e+++iL n++k++l ++++p+sgsl+L++rk +r+frkDG++w+kkkdgktvrE+he+LK g+v+vl+cyYah+e+np+fqrr+yw+Le +
441667 24 EaQNRWLRPLEVIEILRNYQKFRLNPVPPNKPPSGSLFLFDRKTLRFFRKDGHNWRKKKDGKTVREAHERLKAGSVDVLHCYYAHGEDNPNFQRRSYWMLEGA 126
4499*************************************************************************************************** PP
CG-1 106 lekivlvhylev 117
+e+ivlvhy+ev
441667 127 YEHIVLVHYREV 138
**********98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 80.861 | 18 | 144 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 3.2E-74 | 21 | 139 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.2E-48 | 24 | 137 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 2.6E-7 | 439 | 525 | IPR013783 | Immunoglobulin-like fold |
| CDD | cd00102 | 8.03E-4 | 441 | 526 | No hit | No description |
| SuperFamily | SSF81296 | 2.62E-16 | 442 | 525 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 5.2E-6 | 442 | 524 | IPR002909 | IPT domain |
| Gene3D | G3DSA:1.25.40.20 | 2.9E-21 | 614 | 728 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 3.26E-19 | 626 | 726 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 9.24E-19 | 630 | 724 | No hit | No description |
| PROSITE profile | PS50297 | 21.827 | 632 | 736 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 1400 | 632 | 661 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 9.404 | 632 | 664 | IPR002110 | Ankyrin repeat |
| Pfam | PF12796 | 2.2E-9 | 637 | 726 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 10.846 | 665 | 697 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0013 | 665 | 694 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 99 | 704 | 733 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 7.7E-8 | 836 | 889 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.077 | 838 | 860 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.224 | 839 | 868 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0045 | 840 | 859 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.013 | 861 | 883 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.78 | 862 | 886 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0057 | 864 | 883 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 15 | 932 | 954 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.151 | 934 | 962 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 983 aa Download sequence Send to blast |
MYDNRRLRFF PSQPEIDIYQ IIREAQNRWL RPLEVIEILR NYQKFRLNPV PPNKPPSGSL 60 FLFDRKTLRF FRKDGHNWRK KKDGKTVREA HERLKAGSVD VLHCYYAHGE DNPNFQRRSY 120 WMLEGAYEHI VLVHYREVTE GSRSSVYRSM QEAKENASHS RATGQSLPAS NSAISDVEVT 180 GPYKSPEAPV TPIESEGTDS GEEGHFNENN VVEQSSLLQQ VAQSSPVAPP STSVPVAAVP 240 AAAKNLEGVD YDDLLRNPDA YLGQKSIDAQ TWSTLLDNFG GTNTIEKMES TSQSHFLSPG 300 FSNHVYNSIT PTNHPFPPVI PTPDSHNMEV DLRQAQYSAQ DISKKPQTAI PNDASECYKV 360 ALPDVLVEDE GKTSLKKLDS FGRWMSREIG EDSQSSLLSG STDHAYWTLD DHNTFDEISN 420 FTQQIQDVGL GPSVSQDQQF SIVDFSPDWA FASEETKVIV AGNFLKRGAS PVWHCMFGEV 480 EVPAETIHEG VLRCKAPMHS PGRVPLYITL GDRLACSEIR EFEYRTATMK PVAGNPEQLQ 540 VEDEVLEQRF ARLISLNSDE ATKSEEQSDK VQLSKILELT SGLWEDPEPS ESEVGSSTRD 600 TVLQTLLKQQ LQRWLLVKVC DRDKGAAVLD AQGQSALHLA AALGYDWAVN PILAAGVGAN 660 FRDVHGWTGL HWAASRGREK VVSTLLAAGA SPGLVTDPTP QNSSGRTPAD LAASSGHKGM 720 AGLLAEMSLT THLTSLTLKE RNTDEIDSLS AVLAEEKAVE DFSDNQAANG GTDRSLLQGS 780 LRAVRNAARA AALIHASFRQ ESFRRRQEKI GEEIDNEYGM SMNEMKVLAS RNGGANRKEH 840 SAATKIQQKY RGWKGRRDFL LLRQRVVRIQ AHVRGHQVRR KFRKILWTVG ILDKAILRWR 900 RKRGGLRRAS AQTENTDDDD VLKAGRKQKE AQFQKAVTRV QSMVRSHEAQ EQYQRIQEAY 960 LQAVAGDSQA RNENERDETM LG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4gmr_A | 2e-12 | 630 | 724 | 34 | 122 | OR266 DE NOVO PROTEIN |
| 4gmr_B | 2e-12 | 630 | 724 | 34 | 122 | OR266 DE NOVO PROTEIN |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
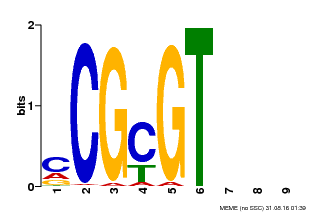
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002971863.1 | 0.0 | calmodulin-binding transcription activator 3 | ||||
| TrEMBL | D8RLU7 | 0.0 | D8RLU7_SELML; Uncharacterized protein | ||||
| STRING | EFJ26780 | 0.0 | (Selaginella moellendorffii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP562 | 15 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 1e-119 | signal responsive 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 441667 |
| Entrez Gene | 9640715 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



