 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Mapoly0074s0081.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Marchantiophyta; Marchantiopsida; Marchantiidae; Marchantiales; Marchantiaceae; Marchantia
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1258aa MW: 141499 Da PI: 7.109 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 177 | 2.5e-55 | 25 | 142 | 3 | 117 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltl..elktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahse 89
+e ++rwl++ e+++iL n++ h+++l ++t+p+sgsl+L++rk +ryfrkDG++w+kkkdgktvrE+he+LK g+v+vl+cyYah+e
Mapoly0074s0081.1.p 25 HEaQTRWLRPVEVCEILRNYRIHNFKLnpVPPTQPQSGSLFLFDRKALRYFRKDGHNWRKKKDGKTVREAHERLKAGSVDVLHCYYAHGE 114
5559*****************9999886689*********************************************************** PP
CG-1 90 enptfqrrcywlLeeelekivlvhylev 117
+n +fqrrcyw+L+ ++e+ivlvhy+ev
Mapoly0074s0081.1.p 115 DNVNFQRRCYWMLDPSYEHIVLVHYREV 142
**************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 82.35 | 20 | 148 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 4.8E-73 | 23 | 143 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 2.9E-50 | 26 | 141 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 5.9E-6 | 679 | 768 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 1.92E-17 | 682 | 768 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 5.9E-6 | 683 | 767 | IPR002909 | IPT domain |
| CDD | cd00102 | 9.12E-6 | 683 | 769 | No hit | No description |
| CDD | cd00204 | 8.38E-15 | 867 | 974 | No hit | No description |
| PROSITE profile | PS50297 | 16.68 | 867 | 974 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.61E-16 | 867 | 976 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.2E-18 | 868 | 978 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 3.1E-6 | 869 | 944 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 5.6E-7 | 915 | 944 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 11.194 | 915 | 947 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 2800 | 954 | 983 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50096 | 6.705 | 1082 | 1108 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 1.33E-5 | 1096 | 1150 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.05 | 1098 | 1120 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.316 | 1099 | 1128 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0075 | 1102 | 1120 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.54 | 1121 | 1143 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.718 | 1123 | 1150 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.004 | 1124 | 1143 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1258 aa Download sequence Send to blast |
MASMTAGRPR NLPPQPEVDV RQIIHEAQTR WLRPVEVCEI LRNYRIHNFK LNPVPPTQPQ 60 SGSLFLFDRK ALRYFRKDGH NWRKKKDGKT VREAHERLKA GSVDVLHCYY AHGEDNVNFQ 120 RRCYWMLDPS YEHIVLVHYR EVTENNRSCI IRTMHDYNST YHPPLASQHQ HVSPTVSVLT 180 NASLPSSVAS PETVEAAGAS LSPELDEAES AEPDEFEDPS LEIEGDQSGL QPSLLEQLGN 240 QPLPGAHSAS SSNQWPSLGR TAKNLGVTDG QFAPLFSHQD NRDFKPSFAD LQQQRPQRMS 300 STSQSVNYPL LQNYPRDVGS IRQQFADMLD SPGPYPSSSQ RSTSSINHSL DLSVLTSMIQ 360 DGNSGHAGNG AERFARPEVN SPPDWAGMLR QVTTPSTRRA LLEGQGLNPG LGQSPHGKDT 420 PPVLGTPISP KGIVDALSPR NLGRAPQTTL EANLRAVTEK EVLKEAEEKA QAQRTRWQTT 480 QLTEPSPVDS VIHRNVDLSG GAQQMSANHF IEPDTLEHLV RSGHSNPDSF STSGSNEKGS 540 WMQQAPQQNE QPVHASQLYQ NGNLYDHQSQ PNQERVQHEN LQPIVPPPHS TGFSFMDHSG 600 LGNGDLFSPT EHYLKDVQSE EGAHREVIES YKKLDSFGRW MTAEIASDIA PYLMVEPGAW 660 DNMQQMQMGG NLDPSPTEQC FNISDFSPEW GSSTEETKVL ITGTFVGSIT PTDHTWACMF 720 GEIEVPAELV EGGALRCKAP KHEPGRVSFY ITRSDRFACS QVREFEYRSA GTTNESTPTL 780 LTGESSKIEE TAEELRYKVR FARMLMATTT LSNCRMNSRT QTFKESEDDD WRQLESMLEA 840 KRPLAEVKEQ LIQLMFKRKL QEWLKEKSLL KDGKRASVHD DQGWGVSHFS AALGYVWSIR 900 PTLNAEIPIN FRDEKGWTAL HWAAYCGREE VVCALLEAGA SPNCATYCNK KYPQGQYPAD 960 LASERGHNGI AAFLSEKALT ARLCGLNLED GTQVDENAAL LAADKAVAQL ARRSSIRSSV 1020 HVEKDHEVLQ TSLLAVRNAT QAAALIQDAN RQASFRRRML HLADDVVDEY GLSQAEVRSM 1080 LAAQKIQKAY RGHQKNKKLH LAATAIQRKF RGWKGRKDFQ TFRQRIIKLQ AHVRGRLVRK 1140 QFRKILWSVS IVEKVVLRWL RRRNGLRGFR SEFIELPTNS DDDDDLMREG RHRSTAAVEK 1200 DVRRVQEMAR EKKAREQYNR LHERISKGEP YNSSYPDHGQ VSRDYMVGQD SVMTTVE* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
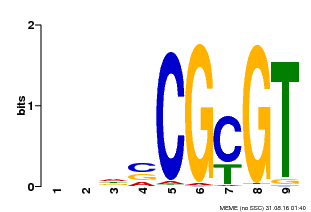
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A176WCI9 | 0.0 | A0A176WCI9_MARPO; Uncharacterized protein | ||||
| TrEMBL | A0A2R6WMN2 | 0.0 | A0A2R6WMN2_MARPO; Uncharacterized protein | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP562 | 15 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 1e-106 | signal responsive 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Mapoly0074s0081.1.p |



