 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MDP0000255517 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Malus
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1122aa MW: 125262 Da PI: 5.8224 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 144.4 | 3e-45 | 74 | 174 | 18 | 118 |
CG-1 18 LenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcywlLeeelekivlvh 113
++ f++ +++ ++ + + gsl L++rk++ryfrkDG++w+kk+dgktv+E+hekLKvg+v+vl+cyYah+e++++fqrrcywlLe++l +iv+vh
MDP0000255517 74 VVVFTHRTFWCATFSAMPGGSLYLFDRKVLRYFRKDGHNWRKKRDGKTVKEAHEKLKVGSVDVLHCYYAHGEDDENFQRRCYWLLEQDLMHIVFVH 169
3445556678888889999***************************************************************************** PP
CG-1 114 ylevk 118
yl+vk
MDP0000255517 170 YLQVK 174
**985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 75.283 | 15 | 179 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 7.1E-65 | 18 | 174 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 2.9E-5 | 21 | 59 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 7.7E-37 | 91 | 172 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 7.8E-5 | 520 | 609 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 1.22E-16 | 523 | 608 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 3.58E-14 | 700 | 825 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 5.7E-15 | 708 | 829 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 14.743 | 709 | 835 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.50E-10 | 709 | 823 | No hit | No description |
| SuperFamily | SSF52540 | 4.51E-8 | 935 | 989 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 1.1 | 938 | 960 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.078 | 939 | 968 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0047 | 941 | 959 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0071 | 961 | 983 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.212 | 962 | 986 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.4E-4 | 964 | 983 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1122 aa Download sequence Send to blast |
MAERGSDAQG RRLDFRQLLV EAQHRWLRPA EICEILSNFH KFHISPEPPN KPXSTANSLA 60 LSVHKTWLRV VVSVVVFTHR TFWCATFSAM PGGSLYLFDR KVLRYFRKDG HNWRKKRDGK 120 TVKEAHEKLK VGSVDVLHCY YAHGEDDENF QRRCYWLLEQ DLMHIVFVHY LQVKGNRANV 180 GGIIENDEVT PDLKRGSPWT SSPSNSNGRT PSGYTDHASP SSTSTSACED VDSGDSRQAS 240 SLFHSVFESP KMGNGPLMDN AEIDPSLHPS SNNHDGQSSI HGVYRPQFEN DQHCFSADST 300 GVIDCQKTLG AVSWEEILEQ CTTGFHTGTS HASMSTTQIG SGEVPVEQQI LGNSFLKEEF 360 GSXMPFQTNW QLPLGDNSLQ LPECSLDQST NLQRTLDANL RNTPDLVSAH LNQRNEQLVQ 420 DDLQAQLTNA ESECLIISSS EHDVPKDGNV NYAFTLRQQL LDREDGLKKV DSFSRWVSKE 480 LEEVDDLQMQ SSSGIPWSTA EYGDVVDDSS LSPSVSQDQL FSIVDFSPKW AYTDSEIEVL 540 VFGTFLVSQK EVTKYSWSCM FGEVEVPAQV LASGVLFCFA PPHSAGQVPF YVTCSNRLAC 600 SEVREFNYQV GSTKGFDMTD IGDGAAHEMH LHLRLESLLS LRPLSPSGQP FGGVKENRNL 660 ISKIISLQEE EDYLAEPTVV NYLLQNEGME HLIKLMKEKL YTWLLYKAIE DGKGPSVLDA 720 EGQGVLHLAA ALGYDWAIKP TVTAGVSINF RDVNGWTALH WAAFCGRQGH QNPFHFILHT 780 VAVLVSLGAD PGALTDPSPE VPLGRTPADL ASVSSHKGIS GFLAESSLTT YLKSLTMNDA 840 KEDGAAEISG TKVVKTFSER IATPVSYGDM PDALSLKDSL TAITNATQAA DRIQQMLRMQ 900 SFERRXLTEY DSDEFGMPDE RAISFITSKS HKGPINRNAH TAAVQIQKKF RGWKKRKEFL 960 IIRQRIVKIQ AHVRGHQVRK QYKAITWSVG ILEKVILRWR RKGTGLRGFR PDAVAKIPDP 1020 QTVASSEDDY DFLKKGRKQT EERLQKALTR VKSMVQYLEG RAQYRRLLNV VEGFGETKVN 1080 DMAVEVSEEK AEGGDEKAEG GDDLIDIDKL LDDDTFMSIA FD |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mdo.9643 | 1e-141 | bud| cell culture| fruit| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, old leaves, petals, sepals, top of carpels, stigmas, stamen filaments, anthers and siliques, but not in pollen. {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:14581622}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
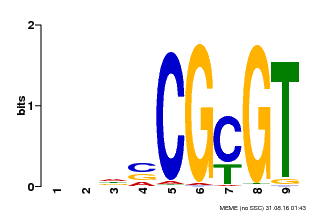
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008358353.2 | 0.0 | calmodulin-binding transcription activator 2-like | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A498J055 | 0.0 | A0A498J055_MALDO; Uncharacterized protein | ||||
| STRING | XP_008358353.1 | 0.0 | (Malus domestica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7406 | 26 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | MDP0000255517 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



