 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LPERR07G22580.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Leersia
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 328aa MW: 33766.9 Da PI: 6.967 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 56.3 | 7.9e-18 | 59 | 107 | 3 | 55 |
AP2 3 ykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+ GVr+++ +gr++AeIrdp++ + r++lg+f+ta+eAa a+++a+++++g
LPERR07G22580.1 59 FLGVRRRP-WGRYAAEIRDPTT---KERHWLGTFDTAQEAALAYDRAALSMKG 107
56******.**********965...5************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00380 | 1.3E-30 | 58 | 121 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 20.547 | 58 | 115 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 2.20E-25 | 58 | 114 | No hit | No description |
| SuperFamily | SSF54171 | 1.05E-19 | 59 | 116 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 5.0E-28 | 59 | 115 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.2E-7 | 59 | 70 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 2.9E-11 | 61 | 107 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.2E-7 | 81 | 97 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010102 | Biological Process | lateral root morphogenesis | ||||
| GO:0010432 | Biological Process | bract development | ||||
| GO:0010451 | Biological Process | floral meristem growth | ||||
| GO:0010582 | Biological Process | floral meristem determinacy | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 328 aa Download sequence Send to blast |
MNTRGGSNGG HSQATLMAFS DQPKPVSHPS PPSSPVSDHR PPSSAGGRGR RRAQEPGRFL 60 GVRRRPWGRY AAEIRDPTTK ERHWLGTFDT AQEAALAYDR AALSMKGAQA RTNFVYTHHH 120 AAAAYNFPQF LAGAFHHHHP SPPAFAAGGG GNSVEMAASA SSHAGTYGGH HVAAGGGGEC 180 STVMSTAMPV MVPVDHFEHH RASTASVNDF LFSGGVYGGG GGGDNSGYLS SVVPESCLRP 240 NGGGGGAADH HDMRRYSDAD AYGMMGIRED VDDLAQMVAG FWGGAADADQ LGVCGFPANG 300 GGGGGGAGDM VAASQGSDSY SPFSFLSH |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wx9_A | 4e-18 | 55 | 114 | 11 | 71 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Required to prevent the formation of axillary meristems within the spikelet meristem and permit the subsequent establishment of floral meristem identity (PubMed:12835399, PubMed:14503923). Mediates the transition from spikelet to floret meristem (PubMed:14503923). Determines the transition from panicle branching to spikelet formation. May specify floral organ identity by regulating the class B genes (Agamous-like genes) MADS6 and MADS17, as well as class E genes MADS1, MADS7 and MADS8 in floral meristem (PubMed:26744119). Possesses transactivation activity (PubMed:12835399). {ECO:0000269|PubMed:12835399, ECO:0000269|PubMed:14503923, ECO:0000269|PubMed:26744119}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00520 | DAP | Transfer from AT5G18560 | Download |
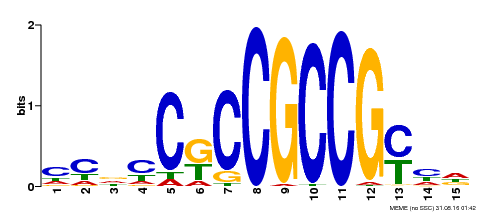
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LPERR07G22580.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY885976 | 4e-76 | AY885976.1 Oryza meridionalis isolate fzp02 frizzy panicle (fzp) gene, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015646388.1 | 1e-108 | ethylene-responsive transcription factor FZP | ||||
| Swissprot | Q8H3Q1 | 1e-109 | FZP_ORYSJ; Ethylene-responsive transcription factor FZP | ||||
| TrEMBL | A0A0D9X2Q8 | 0.0 | A0A0D9X2Q8_9ORYZ; Uncharacterized protein | ||||
| STRING | LPERR07G22580.1 | 0.0 | (Leersia perrieri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP10525 | 28 | 38 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G18560.1 | 8e-38 | ERF family protein | ||||



