 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | kfl00375_0090 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Klebsormidiophyceae; Klebsormidiales; Klebsormidiaceae; Klebsormidium
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1209aa MW: 130236 Da PI: 6.3192 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 156.2 | 6.8e-49 | 34 | 151 | 3 | 117 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltl..elktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfq 95
ke ++rwl++ e+++iL n+++++++l e++ pksg+l L++rk vryfrkDG+ w kkkdgk+v+E+hekLKvg+ e+l+cyYahse++ fq
kfl00375_0090 34 KEaHTRWLRPLEVYDILSNYQHYNFKLnkEAPVLPKSGQLYLFDRKEVRYFRKDGHVWTKKKDGKAVQEAHEKLKVGSLEILHCYYAHSEKDDRFQ 129
55599*************99998887766999**************************************************************** PP
CG-1 96 rrcywlLeeelekivlvhylev 117
rrcyw+L+e++e++vlvhyl+v
kfl00375_0090 130 RRCYWMLDEARERVVLVHYLQV 151
*******************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 68.574 | 29 | 157 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.8E-59 | 32 | 152 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 4.7E-46 | 35 | 150 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 2.94E-10 | 632 | 722 | IPR014756 | Immunoglobulin E-set |
| CDD | cd00102 | 0.00621 | 637 | 723 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 1.7E-21 | 837 | 942 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 4.82E-20 | 839 | 935 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 4.88E-20 | 840 | 940 | No hit | No description |
| SMART | SM00248 | 290 | 842 | 871 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50297 | 22.596 | 842 | 941 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.618 | 842 | 874 | IPR002110 | Ankyrin repeat |
| Pfam | PF12796 | 2.9E-10 | 847 | 932 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 10.846 | 875 | 900 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 2.8E-4 | 875 | 904 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 290 | 909 | 938 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 200 | 987 | 1009 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 4.96E-7 | 1035 | 1091 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 10 | 1043 | 1065 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.309 | 1044 | 1073 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.075 | 1066 | 1088 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 10.109 | 1068 | 1091 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0023 | 1069 | 1088 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 25 | 1145 | 1167 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.389 | 1146 | 1175 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1209 aa Download sequence Send to blast |
MPPPAVGGLH KPFDHYTSLS EIPKPELPIA EILKEAHTRW LRPLEVYDIL SNYQHYNFKL 60 NKEAPVLPKS GQLYLFDRKE VRYFRKDGHV WTKKKDGKAV QEAHEKLKVG SLEILHCYYA 120 HSEKDDRFQR RCYWMLDEAR ERVVLVHYLQ VTEAAHPGDH LQPLGLLPRK ALLHPVKRIP 180 ASESSQGSES EEDRASPNAV LPHPARPRKA LPKRHSFAGG HKGTKRHADD GEGIPADPHR 240 VPKSEPASDG SSQGTGKARR HGEEGNGAQA RGENQETGLT KQGSQDLTSV GGALEAAVRD 300 GVFQMEVDMS RPNGGAALDA LGDVAWQDWQ QQQKVSAAGG YPWDVPAGPP SWPVAPETTA 360 PFPWDPREEP WQGLPRQESA GNLAPPPPLL LPADPGAATG MELWDANNWG GAPPLEGASE 420 AALWLKGEDE RAPAAQMVPV TSADVAPNAV LLNTPSGDWT ASWSGGTPAA ALDLSRAWPD 480 AARTSWPPAG APPSVWVPTQ PPPAHEMPIV HEPGPATGAP SGKPPRPPSL PRLRPLKNLR 540 ALAVPDGAAH SGVGVGSPFL VPESPTVVQV LREMHTDAGS RPGSAAPIPE EAERNEFEER 600 KDEEGAQMEE AGPDERAGGH ARSAGQQLPW FEIQDCSPEW DYSVGGAKVL LVGTWHLPDD 660 SPPGASDDFR ASVTFGGAEV PAEPVGPCLR CTAPAHMPGR VPLCVTFGEN QPASASREFE 720 YRPSPVAYVT RRSAAGGGAL AVEDAVDEDD DAALQVRLAR LLLEGGAGEG GNSDGEEWEE 780 LLRTERVTGC VQDGLLQRVL RHVLHLWLAR AGQGDPLTYD GYAPFGRAGD AAPGGLTGLN 840 EVGLSALHMA AALGYEWAVT ALVDAGCPVD LKDARGRTAL HWAAARGREK VVAQLLTHGA 900 APGMLATKGG CTPADLAATC GHAGIAAFLA ECALNRSLSN MTINEDMAAA SDAAEAGAAA 960 VAVATGSQGV PRGSDAGLEN ALEAVRNAAR AAALIQSAYR RRRRQRQQRA QQGRIPGEGE 1020 RDEAADGMNG MAGGVEEQQK METEHQAATR IQGAFRKFAD RKEFLSFRNK VVRLQAAVRG 1080 YLQRQKYRRM LIGGAYGWRL RSADKRPRSA RYHVDETELP PLPSPLGEAA GEEWRNVGGT 1140 DSNYALDRAV TRVQAMVRSR QARAQYLRLR QASQSMQVEH WQHQHGVMQE PHVRFAEASD 1200 VFDPSVYLV |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
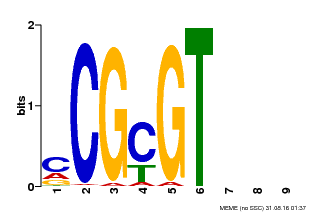
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A1Y1IEB4 | 0.0 | A0A1Y1IEB4_KLENI; Calmodulin-binding protein | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP562 | 15 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G09410.2 | 3e-45 | ethylene induced calmodulin binding protein | ||||



