 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | evm_27.model.AmTr_v1.0_scaffold00001.511 | ||||||||
| Common Name | AMTR_s00001p00272270, LOC18424570 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; basal Magnoliophyta; Amborellales; Amborellaceae; Amborella
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 812aa MW: 88097.2 Da PI: 7.183 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 51.1 | 3.1e-16 | 24 | 68 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+eE++++++a k++G W +I +++g ++t+ q++s+ qk+
evm_27.model.AmTr_v1.0_scaffold00001.511 24 RERWTEEEHDRFLEALKLYGRA-WQRIEEHIG-TKTAVQIRSHAQKF 68
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 8.22E-17 | 18 | 74 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 21.037 | 19 | 73 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 3.9E-17 | 22 | 71 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 5.9E-13 | 23 | 71 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-8 | 24 | 64 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 2.3E-13 | 24 | 67 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.03E-9 | 26 | 69 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043433 | Biological Process | negative regulation of sequence-specific DNA binding transcription factor activity | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 812 aa Download sequence Send to blast |
METYSPGEDL VVKTRKPYTI TKQRERWTEE EHDRFLEALK LYGRAWQRIE EHIGTKTAVQ 60 IRSHAQKFFS KLEREAVIKG LPVGKAHDID IPPPRPKKKP SNPYPRKTGI GSVGPLTGGI 120 KDGSLLMKGP SACCSKAGLN SENSHIIQGS TETASSQRAK ETSIDNENLE VLTLSLGAPC 180 SSVSSASKIS PLVKSKTSTP TSPGAFKEFM PTKKDSGDQT QTDEGSATSK SKKTPKPDNQ 240 SELTELEIGR PDQLNLDYGS AVYVTSTHVP VRDNTDDLKH PRKLELPRKN DTQATQNYPR 300 HVPVHVVDGT LNKCMHSSSS NPTNSVSTVH QMGVHGNPRL IATPSLSVTS DSAASTMCPQ 360 FPMFHPSFTP FHLNPEAYKL LAPSAMNLSC VLSSYIIANL LQNPAAHAAA TLAASFWPGL 420 DFETPLDATT AEILRSSAPL NYVDVSPPNI AAVVAATVAA ASAWWASHGP LPLSHPHPHP 480 HHTCFFAPSA TLVPVAEPSP VSEEKKEEKN KESMDQDGPP FKQPQKAQAR DADFSPNLQA 540 RIPVSTMASS SDLEDSEGVG SDNSKPKVSE HEQKLLAVEV VNGKPRKQLD RSSCGSNASS 600 SSDIETDNLE KNDEGKEASP VAEFCYSGNE FGRRSRTAGA ISDSWKEVSE GGRLAFQALF 660 SREVLPQSFS PPHNQKDKTS NATHIDGEVK RKDMSKSSDS KEGNVDLNTA TWVEDHCTNE 720 DSSDTTRGNR PQTGSSVSEE EKGNGKTDTL NGKLQACRMG FEPYKRCSVE AKECRLVDGT 780 REKGLKRIRL EHESNSPTSV GSFLKRAHCP S* |
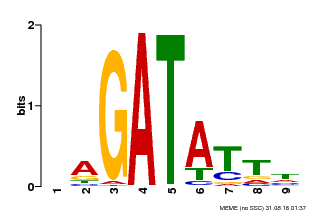
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006829218.1 | 0.0 | protein LHY | ||||
| Refseq | XP_020532316.1 | 0.0 | protein LHY | ||||
| TrEMBL | W1NMV4 | 0.0 | W1NMV4_AMBTC; Uncharacterized protein | ||||
| STRING | ERM96634 | 0.0 | (Amborella trichopoda) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP9903 | 5 | 10 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46830.1 | 3e-45 | circadian clock associated 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | evm_27.model.AmTr_v1.0_scaffold00001.511 |
| Entrez Gene | 18424570 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



