- Riechmann JL, et al.
Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes.
Science, 2000. 290(5499): p. 2105-10
[PMID:11118137] - Stracke R,Werber M,Weisshaar B
The R2R3-MYB gene family in Arabidopsis thaliana.
Curr. Opin. Plant Biol., 2001. 4(5): p. 447-56
[PMID:11597504] - Kersten B, et al.
Generation of Arabidopsis protein chips for antibody and serum screening.
Plant Mol. Biol., 2003. 52(5): p. 999-1010
[PMID:14558660] - Yamada K, et al.
Empirical analysis of transcriptional activity in the Arabidopsis genome.
Science, 2003. 302(5646): p. 842-6
[PMID:14593172] - Leonhardt N, et al.
Microarray expression analyses of Arabidopsis guard cells and isolation of a recessive abscisic acid hypersensitive protein phosphatase 2C mutant.
Plant Cell, 2004. 16(3): p. 596-615
[PMID:14973164] - Ko JH,Han KH,Park S,Yang J
Plant body weight-induced secondary growth in Arabidopsis and its transcription phenotype revealed by whole-transcriptome profiling.
Plant Physiol., 2004. 135(2): p. 1069-83
[PMID:15194820] - Reymond P, et al.
A conserved transcript pattern in response to a specialist and a generalist herbivore.
Plant Cell, 2004. 16(11): p. 3132-47
[PMID:15494554] - Devoto A, et al.
Expression profiling reveals COI1 to be a key regulator of genes involved in wound- and methyl jasmonate-induced secondary metabolism, defence, and hormone interactions.
Plant Mol. Biol., 2005. 58(4): p. 497-513
[PMID:16021335] - Lee BH,Henderson DA,Zhu JK
The Arabidopsis cold-responsive transcriptome and its regulation by ICE1.
Plant Cell, 2005. 17(11): p. 3155-75
[PMID:16214899] - Yanhui C, et al.
The MYB transcription factor superfamily of Arabidopsis: expression analysis and phylogenetic comparison with the rice MYB family.
Plant Mol. Biol., 2006. 60(1): p. 107-24
[PMID:16463103] - Ma S,Bohnert HJ
Integration of Arabidopsis thaliana stress-related transcript profiles, promoter structures, and cell-specific expression.
Genome Biol., 2007. 8(4): p. R49
[PMID:17408486] - Shin R, et al.
The Arabidopsis transcription factor MYB77 modulates auxin signal transduction.
Plant Cell, 2007. 19(8): p. 2440-53
[PMID:17675404] - Libault M,Wan J,Czechowski T,Udvardi M,Stacey G
Identification of 118 Arabidopsis transcription factor and 30 ubiquitin-ligase genes responding to chitin, a plant-defense elicitor.
Mol. Plant Microbe Interact., 2007. 20(8): p. 900-11
[PMID:17722694] - Wang Z, et al.
Identification and characterization of COI1-dependent transcription factor genes involved in JA-mediated response to wounding in Arabidopsis plants.
Plant Cell Rep., 2008. 27(1): p. 125-35
[PMID:17786451] - Jung C, et al.
Overexpression of AtMYB44 enhances stomatal closure to confer abiotic stress tolerance in transgenic Arabidopsis.
Plant Physiol., 2008. 146(2): p. 623-35
[PMID:18162593] - Sreenivasulu N, et al.
Barley grain maturation and germination: metabolic pathway and regulatory network commonalities and differences highlighted by new MapMan/PageMan profiling tools.
Plant Physiol., 2008. 146(4): p. 1738-58
[PMID:18281415] - Huang D,Wu W,Abrams SR,Cutler AJ
The relationship of drought-related gene expression in Arabidopsis thaliana to hormonal and environmental factors.
J. Exp. Bot., 2008. 59(11): p. 2991-3007
[PMID:18552355] - Pitzschke A,Djamei A,Teige M,Hirt H
VIP1 response elements mediate mitogen-activated protein kinase 3-induced stress gene expression.
Proc. Natl. Acad. Sci. U.S.A., 2009. 106(43): p. 18414-9
[PMID:19820165] - Jung C, et al.
Non-specific phytohormonal induction of AtMYB44 and suppression of jasmonate-responsive gene activation in Arabidopsis thaliana.
Mol. Cells, 2010. 29(1): p. 71-6
[PMID:20016937] - Liu R, et al.
Thirty-seven transcription factor genes differentially respond to a harpin protein and affect resistance to the green peach aphid in Arabidopsis.
J. Biosci., 2010. 35(3): p. 435-50
[PMID:20826953] - Liu R, et al.
Transcription factor AtMYB44 regulates induced expression of the ETHYLENE INSENSITIVE2 gene in Arabidopsis responding to a harpin protein.
Mol. Plant Microbe Interact., 2011. 24(3): p. 377-89
[PMID:21117868] - L
HrpN Ea-induced deterrent effect on phloem feeding of the green peach aphid Myzus persicae requires AtGSL5 and AtMYB44 genes in Arabidopsis thaliana.
J. Biosci., 2011. 36(1): p. 123-37
[PMID:21451254] - Causier B,Ashworth M,Guo W,Davies B
The TOPLESS interactome: a framework for gene repression in Arabidopsis.
Plant Physiol., 2012. 158(1): p. 423-38
[PMID:22065421] - Nguyen XC, et al.
Phosphorylation of the transcriptional regulator MYB44 by mitogen activated protein kinase regulates Arabidopsis seed germination.
Biochem. Biophys. Res. Commun., 2012. 423(4): p. 703-8
[PMID:22704933] - Shim JS, et al.
AtMYB44 regulates WRKY70 expression and modulates antagonistic interaction between salicylic acid and jasmonic acid signaling.
Plant J., 2013. 73(3): p. 483-95
[PMID:23067202] - Jung C, et al.
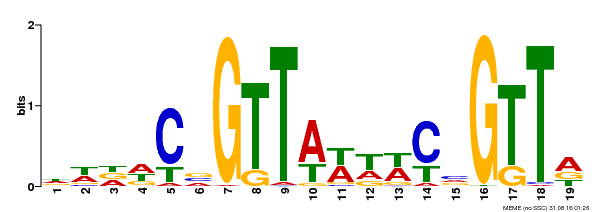
Quadruple 9-mer-based protein binding microarray analysis confirms AACnG as the consensus nucleotide sequence sufficient for the specific binding of AtMYB44.
Mol. Cells, 2012. 34(6): p. 531-7
[PMID:23161171] - Persak H,Pitzschke A
Tight interconnection and multi-level control of Arabidopsis MYB44 in MAPK cascade signalling.
PLoS ONE, 2013. 8(2): p. e57547
[PMID:23437396] - Shim JS,Choi YD
Direct regulation of WRKY70 by AtMYB44 in plant defense responses.
Plant Signal Behav, 2013. 8(6): p. e20783
[PMID:23603962] - L
AtMYB44 regulates resistance to the green peach aphid and diamondback moth by activating EIN2-affected defences in Arabidopsis.
Plant Biol (Stuttg), 2013. 15(5): p. 841-50
[PMID:23656500] - Li C,Chang PP,Ghebremariam KM,Qin L,Liang Y
Overexpression of tomato SpMPK3 gene in Arabidopsis enhances the osmotic tolerance.
Biochem. Biophys. Res. Commun., 2014. 443(2): p. 357-62
[PMID:24275141] - Jaradat MR,Feurtado JA,Huang D,Lu Y,Cutler AJ
Multiple roles of the transcription factor AtMYBR1/AtMYB44 in ABA signaling, stress responses, and leaf senescence.
BMC Plant Biol., 2013. 13: p. 192
[PMID:24286353] - Ding Y, et al.
Four distinct types of dehydration stress memory genes in Arabidopsis thaliana.
BMC Plant Biol., 2013. 13: p. 229
[PMID:24377444] - Persak H,Pitzschke A
Dominant repression by Arabidopsis transcription factor MYB44 causes oxidative damage and hypersensitivity to abiotic stress.
Int J Mol Sci, 2014. 15(2): p. 2517-37
[PMID:24531138] - Li D, et al.
Arabidopsis ABA receptor RCAR1/PYL9 interacts with an R2R3-type MYB transcription factor, AtMYB44.
Int J Mol Sci, 2014. 15(5): p. 8473-90
[PMID:24828206] - Zhao Y, et al.
The ABA receptor PYL8 promotes lateral root growth by enhancing MYB77-dependent transcription of auxin-responsive genes.
Sci Signal, 2014. 7(328): p. ra53
[PMID:24894996] - Xu DB, et al.
A G-protein β subunit, AGB1, negatively regulates the ABA response and drought tolerance by down-regulating AtMPK6-related pathway in Arabidopsis.
PLoS ONE, 2015. 10(1): p. e0116385
[PMID:25635681] - Jin J, et al.
An Arabidopsis Transcriptional Regulatory Map Reveals Distinct Functional and Evolutionary Features of Novel Transcription Factors.
Mol. Biol. Evol., 2015. 32(7): p. 1767-73
[PMID:25750178] - Hieno A, et al.
Possible Involvement of MYB44-Mediated Stomatal Regulation in Systemic Resistance Induced by Penicillium simplicissimum GP17-2 in Arabidopsis.
Microbes Environ., 2016. 31(2): p. 154-9
[PMID:27301421] - Zhao Q, et al.
AtMYB44 Positively Regulates the Enhanced Elongation of Primary Roots Induced by N-3-Oxo-Hexanoyl-Homoserine Lactone in Arabidopsis thaliana.
Mol. Plant Microbe Interact., 2016. 29(10): p. 774-785
[PMID:27604593] - Song L, et al.
A transcription factor hierarchy defines an environmental stress response network.
Science, 2017.
[PMID:27811239] - Nguyen NH,Cheong JJ
H2A.Z-containing nucleosomes are evicted to activate AtMYB44 transcription in response to salt stress.
Biochem. Biophys. Res. Commun., 2018. 499(4): p. 1039-1043
[PMID:29649476] - Nguyen NH,Cheong JJ
AtMYB44 interacts with TOPLESS-RELATED corepressors to suppress protein phosphatase 2C gene transcription.
Biochem. Biophys. Res. Commun., 2018. 507(1-4): p. 437-442
[PMID:30448055] - Kirik V,K
Two novel MYB homologues with changed expression in late embryogenesis-defective Arabidopsis mutants.
Plant Mol. Biol., 1998. 37(5): p. 819-27
[PMID:9678577]
|