| Signature Domain? help Back to Top |
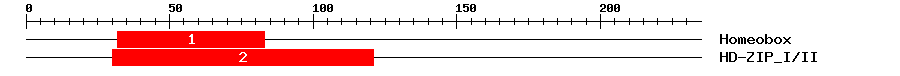 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | Homeobox | 54.8 | 1.6e-17 | 32 | 83 | 5 | 56 |
SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 5 ttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+f++eq++ Le Fe++ + +++ + A++lgL+ rqV +WFqN+Ra++k
AT3G61890.1 32 KRFSEEQIKSLELIFESETRLEPRKKVQVARELGLQPRQVAIWFQNKRARWK 83
68*************************************************9 PP
|
| 2 | HD-ZIP_I/II | 117.9 | 5.5e-38 | 30 | 121 | 2 | 93 |
HD-ZIP_I/II 2 kkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
+++r+s+eq+k+LE Fe+e++Lep++Kv++areLglqprqva+WFqn+RAR+ktkqlEk+y++L+++y++l+++ e ++ke+++L +el++
AT3G61890.1 30 NQKRFSEEQIKSLELIFESETRLEPRKKVQVARELGLQPRQVAIWFQNKRARWKTKQLEKEYNTLRANYNNLASQFEIMKKEKQSLVSELQR 121
689*************************************************************************************9986 PP
|
| Publications
? help Back to Top |
- Riechmann JL, et al.
Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes.
Science, 2000. 290(5499): p. 2105-10
[PMID:11118137] - Johannesson H,Wang Y,Engström P
DNA-binding and dimerization preferences of Arabidopsis homeodomain-leucine zipper transcription factors in vitro.
Plant Mol. Biol., 2001. 45(1): p. 63-73
[PMID:11247607] - Lee YH, et al.
Structure and expression of the Arabidopsis thaliana homeobox gene Athb-12.
Biochem. Biophys. Res. Commun., 2001. 284(1): p. 133-41
[PMID:11374882] - Bray EA
Classification of genes differentially expressed during water-deficit stress in Arabidopsis thaliana: an analysis using microarray and differential expression data.
Ann. Bot., 2002. 89 Spec No: p. 803-11
[PMID:12102506] - Yamada K, et al.
Empirical analysis of transcriptional activity in the Arabidopsis genome.
Science, 2003. 302(5646): p. 842-6
[PMID:14593172] - Shin D, et al.
Athb-12, a homeobox-leucine zipper domain protein from Arabidopsis thaliana, increases salt tolerance in yeast by regulating sodium exclusion.
Biochem. Biophys. Res. Commun., 2004. 323(2): p. 534-40
[PMID:15369784] - Olsson AS,Engström P,Söderman E
The homeobox genes ATHB12 and ATHB7 encode potential regulators of growth in response to water deficit in Arabidopsis.
Plant Mol. Biol., 2004. 55(5): p. 663-77
[PMID:15604708] - Henriksson E, et al.
Homeodomain leucine zipper class I genes in Arabidopsis. Expression patterns and phylogenetic relationships.
Plant Physiol., 2005. 139(1): p. 509-18
[PMID:16055682] - Moes D,Himmelbach A,Korte A,Haberer G,Grill E
Nuclear localization of the mutant protein phosphatase abi1 is required for insensitivity towards ABA responses in Arabidopsis.
Plant J., 2008. 54(5): p. 806-19
[PMID:18298671] - Huang D,Wu W,Abrams SR,Cutler AJ
The relationship of drought-related gene expression in Arabidopsis thaliana to hormonal and environmental factors.
J. Exp. Bot., 2008. 59(11): p. 2991-3007
[PMID:18552355] - Ascencio-Ib
Global analysis of Arabidopsis gene expression uncovers a complex array of changes impacting pathogen response and cell cycle during geminivirus infection.
Plant Physiol., 2008. 148(1): p. 436-54
[PMID:18650403] - Son O, et al.
ATHB12, an ABA-inducible homeodomain-leucine zipper (HD-Zip) protein of Arabidopsis, negatively regulates the growth of the inflorescence stem by decreasing the expression of a gibberellin 20-oxidase gene.
Plant Cell Physiol., 2010. 51(9): p. 1537-47
[PMID:20668225] - Park J, et al.
The Arabidopsis thaliana homeobox gene ATHB12 is involved in symptom development caused by geminivirus infection.
PLoS ONE, 2011. 6(5): p. e20054
[PMID:21625602] - Vald
The homeodomain-leucine zipper (HD-Zip) class I transcription factors ATHB7 and ATHB12 modulate abscisic acid signalling by regulating protein phosphatase 2C and abscisic acid receptor gene activities.
Plant Mol. Biol., 2012. 80(4-5): p. 405-18
[PMID:22968620] - Ding Y, et al.
Four distinct types of dehydration stress memory genes in Arabidopsis thaliana.
BMC Plant Biol., 2013. 13: p. 229
[PMID:24377444] - Capella M,R
Plant homeodomain-leucine zipper I transcription factors exhibit different functional AHA motifs that selectively interact with TBP or/and TFIIB.
Plant Cell Rep., 2014. 33(6): p. 955-67
[PMID:24531799] - Lumba S, et al.
A mesoscale abscisic acid hormone interactome reveals a dynamic signaling landscape in Arabidopsis.
Dev. Cell, 2014. 29(3): p. 360-72
[PMID:24823379] - R
Arabidopsis AtHB7 and AtHB12 evolved divergently to fine tune processes associated with growth and responses to water stress.
BMC Plant Biol., 2014. 14: p. 150
[PMID:24884528] - Hur YS, et al.
Arabidopsis thaliana homeobox 12 (ATHB12), a homeodomain-leucine zipper protein, regulates leaf growth by promoting cell expansion and endoreduplication.
New Phytol., 2015. 205(1): p. 316-28
[PMID:25187356] - Jin J, et al.
An Arabidopsis Transcriptional Regulatory Map Reveals Distinct Functional and Evolutionary Features of Novel Transcription Factors.
Mol. Biol. Evol., 2015. 32(7): p. 1767-73
[PMID:25750178] - Liu C,Wang B,Li Z,Peng Z,Zhang J
TsNAC1 Is a Key Transcription Factor in Abiotic Stress Resistance and Growth.
Plant Physiol., 2018. 176(1): p. 742-756
[PMID:29122985] - Huang KC,Lin WC,Cheng WH
Salt hypersensitive mutant 9, a nucleolar APUM23 protein, is essential for salt sensitivity in association with the ABA signaling pathway in Arabidopsis.
BMC Plant Biol., 2018. 18(1): p. 40
[PMID:29490615] - Lee YH,Chun JY
A new homeodomain-leucine zipper gene from Arabidopsis thaliana induced by water stress and abscisic acid treatment.
Plant Mol. Biol., 1998. 37(2): p. 377-84
[PMID:9617808]
|