 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | mrna25646.1-v1.0-hybrid | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 962aa MW: 105505 Da PI: 6.9712 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58.2 | 1.4e-18 | 14 | 72 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++++++ps +r++L +++ +++ +q+kvWFqNrR +ek+
mrna25646.1-v1.0-hybrid 14 KYVRYTPEQVEALERLYHECPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 72
5679*****************************************************97 PP
| |||||||
| 2 | START | 165.8 | 3e-52 | 158 | 366 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECT CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetleviss 87
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v +++ v+e+l+d + W ++++ +++l+v+ +
mrna25646.1-v1.0-hybrid 158 IAEETLAEFLSKATGTAVEWVQMPGMKPGPDSIGIVAISHGCPGVAARACGLVGLEPT-RVAEILKDLPSWLRDCRTVDVLNVLPT 242
7899******************************************************.**************************9 PP
T..EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEE CS
START 88 g..galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwv 167
+ g+++l +++l+a+++l+p df+++Ry+ l++g++v++ +S+ ++q+ p+ +++vRae+lpSg+li+p+++g+s +++v
mrna25646.1-v1.0-hybrid 243 AngGTIELLYMQLYAPTTLAPaCDFWMLRYTSVLEDGSLVVCVRSLKNTQNGPSmppVQHFVRAEMLPSGYLIRPCEGGGSIIHIV 328
999*************************************************9988899*************************** PP
E-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 168 ehvdlkgrlphwllrslvksglaegaktwvatlqrqce 205
+h+dl+ ++++lr+l++s+++ ++k+++a+l+++++
mrna25646.1-v1.0-hybrid 329 DHMDLEPCGVPEVLRPLYESSAVLAQKMTMAALRQLRQ 366
**********************************9876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.483 | 9 | 73 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.2E-15 | 11 | 77 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.15E-16 | 14 | 74 | No hit | No description |
| SuperFamily | SSF46689 | 6.84E-17 | 14 | 77 | IPR009057 | Homeodomain-like |
| Pfam | PF00046 | 3.5E-16 | 15 | 72 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.9E-18 | 16 | 72 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 1.36E-6 | 66 | 105 | No hit | No description |
| PROSITE profile | PS50848 | 22.472 | 148 | 376 | IPR002913 | START domain |
| CDD | cd08875 | 1.37E-74 | 152 | 368 | No hit | No description |
| SMART | SM00234 | 6.0E-40 | 157 | 367 | IPR002913 | START domain |
| Gene3D | G3DSA:3.30.530.20 | 1.5E-19 | 157 | 363 | IPR023393 | START-like domain |
| SuperFamily | SSF55961 | 1.92E-35 | 157 | 368 | No hit | No description |
| Pfam | PF01852 | 7.6E-50 | 158 | 366 | IPR002913 | START domain |
| Pfam | PF08670 | 1.5E-46 | 693 | 830 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0007134 | developmental stage | sporophyte vegetative stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 962 aa Download sequence Send to blast |
MSCKDGKGVM DNGKYVRYTP EQVEALERLY HECPKPSSIR RQQLIRECPI LSNIEPKQIK 60 VWFQNRRCRE KQRKEASRLQ AVNRKLTAMN KLLMEENDRL QKQVSHLVYE NGYFRQHTQT 120 TTLATKDTSC ESVVTSGQHH LTPQHPPRDA SPAGLLSIAE ETLAEFLSKA TGTAVEWVQM 180 PGMKPGPDSI GIVAISHGCP GVAARACGLV GLEPTRVAEI LKDLPSWLRD CRTVDVLNVL 240 PTANGGTIEL LYMQLYAPTT LAPACDFWML RYTSVLEDGS LVVCVRSLKN TQNGPSMPPV 300 QHFVRAEMLP SGYLIRPCEG GGSIIHIVDH MDLEPCGVPE VLRPLYESSA VLAQKMTMAA 360 LRQLRQIAHE VSQSSVTGWG RRPAALRALS QRLSRGFNEA LNGFTDEGWS LMGNDGMDDV 420 TILINSSPDK LMGLNLSFGN GFPAVSNAVL CAKASMLLQN VPPAILLRFL REHRSEWADN 480 NIDAYSAAAV KVGPCSLAGS RVGSFGGQVI LPLAHTIEHE EFLEVIKLEG IGHSPDDPMM 540 QTREMFLLQL CSGMDETAVG SCAELIFAPI DASFADDAPL LPSGFRIIPL DSGKEASSPN 600 RTLDLASALE IGPTGNRASS EYSANAGCVR SVMTIAFEFA CESHMQEHVA SMARQYVRSI 660 ISSVQRVALA LSPSHLSSHA GLRSPLGTPE AQTLARWICN SYRGYLGVEL LKSCNEGSES 720 NLKSLWHHSD AIMCCSLKAV PVFTFANQAG LDMLETTLVA LQDIALEKIF DDHGRKTLFS 780 EFPQIMQQGF ACLQGGICLS SMGRPVSYER AVAWKVLNEE ETANCICFFI FMGSLFSQVW 840 GLRMCLVFLD NSKSLFSHVK VSGQAAEGNV PGRVQRVSHL ALVRLRRCWP MAKPECEECG 900 RTGNQVIGAI GDGWVGCHRW LDSDLLSVIR VSLLRLSTPH YLTAAGGDGD IIVSKHKWRV 960 W* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
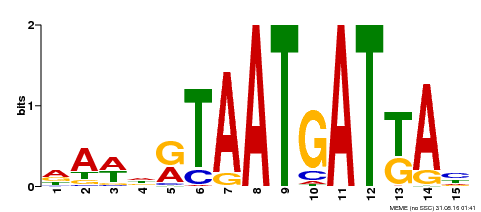
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | mrna25646.1-v1.0-hybrid |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KJ000553 | 0.0 | KJ000553.1 Prunus persica homeobox protein 15 gene, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004302866.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A2P6RXA7 | 0.0 | A0A2P6RXA7_ROSCH; Putative transcription factor & lipid binding Homobox-WOX family | ||||
| STRING | XP_004302866.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1946 | 33 | 87 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | mrna25646.1-v1.0-hybrid |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



