 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | mrna24494.1-v1.0-hybrid | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | MIKC_MADS | ||||||||
| Protein Properties | Length: 263aa MW: 30268.4 Da PI: 10.057 | ||||||||
| Description | MIKC_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 98.4 | 2.9e-31 | 29 | 78 | 1 | 50 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeys 50
krien++nrqvtf+kRrng+lKKA+ELSvLCdaeva+i+fs++g+lyey+
mrna24494.1-v1.0-hybrid 29 KRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSTRGRLYEYA 78
79***********************************************8 PP
| |||||||
| 2 | K-box | 114.2 | 1.3e-37 | 96 | 192 | 4 | 100 |
K-box 4 ssgksleeakaeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqe 89
s++ s++ea+ + +qqe++kL+++i+++q+++Rh+lGe L++L++keL++Le +Lek++++iRskKne+l+++ie++qk+e elq+
mrna24494.1-v1.0-hybrid 96 SNTGSVTEANVQFYQQEASKLRRQIREIQNSNRHILGEALSTLNVKELKNLEGRLEKGISRIRSKKNEMLFAEIEYMQKREIELQN 181
334449******************************************************************************** PP
K-box 90 enkaLrkklee 100
+n+ Lr+k++e
mrna24494.1-v1.0-hybrid 182 HNNFLRAKIAE 192
********986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50066 | 33.501 | 21 | 81 | IPR002100 | Transcription factor, MADS-box |
| SMART | SM00432 | 2.9E-40 | 21 | 80 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 2.88E-33 | 22 | 108 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 1.42E-44 | 22 | 96 | No hit | No description |
| PRINTS | PR00404 | 9.6E-33 | 23 | 43 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 23 | 77 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 2.2E-26 | 30 | 77 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 9.6E-33 | 43 | 58 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 9.6E-33 | 58 | 79 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 1.6E-27 | 104 | 190 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 15.125 | 106 | 196 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0090376 | Biological Process | seed trichome differentiation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009001 | anatomy | fruit | ||||
| PO:0007010 | developmental stage | whole plant fruit ripening stage | ||||
| PO:0007027 | developmental stage | whole plant fruit formation stage 70% to final size | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 263 aa Download sequence Send to blast |
MEIPKQITPA DDPESSSQKK LGRGKIEIKR IENTTNRQVT FCKRRNGLLK KAYELSVLCD 60 AEVALIVFST RGRLYEYANN SVRATIERYK KACDSSNTGS VTEANVQFYQ QEASKLRRQI 120 REIQNSNRHI LGEALSTLNV KELKNLEGRL EKGISRIRSK KNEMLFAEIE YMQKREIELQ 180 NHNNFLRAKI AENDRAQQQQ ANMLPGTSSA YEQPMPPPQS YDRSFLPVII ESNHNYNRQG 240 QNLTPLQLVF DTRLWASFRL SL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1tqe_P | 7e-20 | 21 | 89 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 1tqe_Q | 7e-20 | 21 | 89 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 1tqe_R | 7e-20 | 21 | 89 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 1tqe_S | 7e-20 | 21 | 89 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 5f28_A | 7e-20 | 21 | 89 | 1 | 69 | MEF2C |
| 5f28_B | 7e-20 | 21 | 89 | 1 | 69 | MEF2C |
| 5f28_C | 7e-20 | 21 | 89 | 1 | 69 | MEF2C |
| 5f28_D | 7e-20 | 21 | 89 | 1 | 69 | MEF2C |
| 6c9l_A | 7e-20 | 21 | 89 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_B | 7e-20 | 21 | 89 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_C | 7e-20 | 21 | 89 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_D | 7e-20 | 21 | 89 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_E | 7e-20 | 21 | 89 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_F | 7e-20 | 21 | 89 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in flower development. {ECO:0000250|UniProtKB:Q0HA25}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00093 | SELEX | Transfer from AT3G58780 | Download |
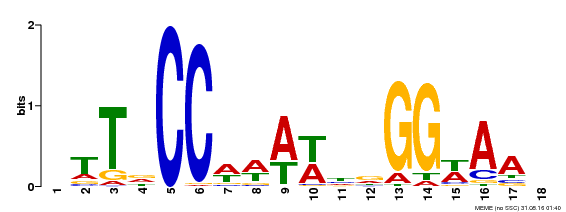
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | mrna24494.1-v1.0-hybrid |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KC676787 | 0.0 | KC676787.1 Fragaria x ananassa SHATTERPROOF-like protein (SHP) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004309724.1 | 0.0 | PREDICTED: floral homeotic protein AGAMOUS isoform X1 | ||||
| Refseq | XP_011471013.1 | 0.0 | PREDICTED: floral homeotic protein AGAMOUS isoform X2 | ||||
| Swissprot | Q93XH4 | 1e-122 | MADS1_VITVI; Agamous-like MADS-box protein MADS1 | ||||
| TrEMBL | A0A2P6S026 | 1e-178 | A0A2P6S026_ROSCH; Putative transcription factor MADS-MIKC family | ||||
| TrEMBL | T2AAD6 | 1e-178 | T2AAD6_FRAAN; SHATTERPROOF-like protein | ||||
| STRING | XP_004309724.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF684 | 29 | 102 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G58780.1 | 1e-117 | MIKC_MADS family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | mrna24494.1-v1.0-hybrid |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



