 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | maker-scaffold00491-augustus-gene-1.11-mRNA-1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Fagaceae; Castanea
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 363aa MW: 40986 Da PI: 6.1986 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.9 | 3.9e-32 | 75 | 128 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtpeLH +Fv+ave+LGG+e+AtPk +l+lm++kgL+++hvkSHLQ+YR+
maker-scaffold00491-augustus-gene-1.11-mRNA-1 75 PRLRWTPELHLCFVHAVERLGGQERATPKLVLQLMNIKGLSIAHVKSHLQMYRS 128
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 12.281 | 71 | 131 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.1E-30 | 71 | 129 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 5.55E-15 | 73 | 129 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 9.8E-23 | 75 | 130 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.1E-9 | 76 | 127 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 363 aa Download sequence Send to blast |
MKSGDDENTQ ASKTSSPSNK SDDDDDDDED EGDEEVDETG KPKNAGSSSN ITVENSSEKK 60 SASGSVRQYV RSKMPRLRWT PELHLCFVHA VERLGGQERA TPKLVLQLMN IKGLSIAHVK 120 SHLQMYRSKK MDDHNRVPNQ GLFLENGDHN IYNLSQLPML QNFNQRPSGL RYGDASWRGH 180 DKQIYGPYMG RAALDRTRNG LYGSVTEKIF ASNNTSTSAN FNLHKRDSSF NGQATWPIRH 240 QTLNEFKWFQ GSKQIPLKQS SLETNLITQL QERGTDDQGI YLNNSGSPRR KIQEAQNTMK 300 RKALDNTDHG GLDLNLSLKA ASNNNEFEKG LDGDDKVEGR RWSFEAFEDD EYSGSDFMNE 360 TIL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 6e-19 | 76 | 130 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 6e-19 | 76 | 130 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 6e-19 | 76 | 130 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 6e-19 | 76 | 130 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00301 | DAP | Transfer from AT2G40260 | Download |
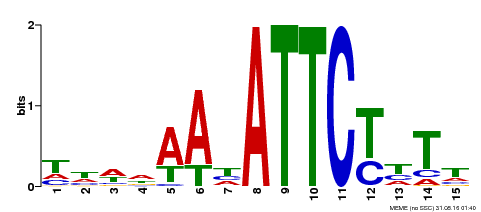
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018821715.1 | 1e-116 | PREDICTED: uncharacterized protein LOC108991786 | ||||
| TrEMBL | A0A2I4EQN1 | 1e-115 | A0A2I4EQN1_JUGRE; uncharacterized protein LOC108991786 | ||||
| STRING | POPTR_0010s19310.1 | 1e-105 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2378 | 34 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40260.1 | 7e-51 | G2-like family protein | ||||



