|
Plant Transcription
Factor Database
|
Transcription Factor Information
|
Basic
Information? help
Back to Top |
| TF ID |
maker-scaffold00401-snap-gene-0.14-mRNA-1 |
| Organism |
|
| Taxonomic ID |
|
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Fagaceae; Castanea
|
| Family |
HD-ZIP |
| Protein Properties |
Length: 322aa MW: 36227.3 Da PI: 4.8212 |
| Description |
HD-ZIP family protein |
| Gene Model |
| Gene Model ID |
Type |
Source |
Coding Sequence |
| maker-scaffold00401-snap-gene-0.14-mRNA-1 | genome | THGP | View Nucleic Acid |
|
| Signature Domain? help Back to Top |
 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | Homeobox | 59.4 | 5.8e-19 | 57 | 110 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe ++++ e++ +LA++lgL+ rqV vWFqNrRa++k
maker-scaffold00401-snap-gene-0.14-mRNA-1 57 KKRRLSVEQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWK 110
56689************************************************9 PP
|
| 2 | HD-ZIP_I/II | 129.8 | 1.1e-41 | 56 | 148 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkr 68
ekkrrls eqvk+LE++Fe e+kLeperKv+la+eLglqprqvavWFqnrRAR+ktkqlE+dy +Lk+
maker-scaffold00401-snap-gene-0.14-mRNA-1 56 EKKRRLSVEQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWKTKQLERDYGVLKA 123
69****************************************************************** PP
HD-ZIP_I/II 69 aydalkeenerLekeveeLreelke 93
+y++lk + ++L++++e+L +e+ke
maker-scaffold00401-snap-gene-0.14-mRNA-1 124 SYETLKLNCDNLQHDNEALLKEIKE 148
********************99987 PP
|
| Gene Ontology ? help Back to Top |
| GO Term |
GO Category |
GO Description |
| GO:0009414 | Biological Process | response to water deprivation |
| GO:0009788 | Biological Process | negative regulation of abscisic acid-activated signaling pathway |
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated |
| GO:0005634 | Cellular Component | nucleus |
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding |
| GO:0043565 | Molecular Function | sequence-specific DNA binding |
| Sequence ? help Back to Top |
| Protein Sequence Length: 322 aa
Download sequence Send
to blast |
MKRLGSSDSL GALMSICPTT DEHSPRTNHV YSREFQSMLD GLDEEGCVEE SGHIAEKKRR 60
LSVEQVKALE KNFEVENKLE PERKVKLAQE LGLQPRQVAV WFQNRRARWK TKQLERDYGV 120
LKASYETLKL NCDNLQHDNE ALLKEIKELK AKLHEENTES VNLSVKEEIV VAESDNKAME 180
QSKPAPDSPP LPASDSKDLN FKCFNHNNGV GGATLFTVDF KDGSSDSDSS AILNEDNSPN 240
AAISSSGVLQ NQLLMSPSQA ASSSSLKFNS SSPMGCFQFQ KTYQPQFVKM EEHNFFSGEE 300
ACNFFSDEQP PTLQWYCSEQ WN
|
| Nucleic Localization
Signal ? help
Back to Top |
 |
| No. |
Start |
End |
Sequence |
| 1 | 104 | 112 | RRARWKTKQ |
| Functional Description ? help
Back to Top |
| Source |
Description |
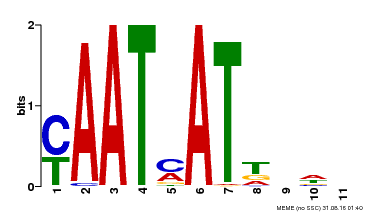
| UniProt | Transcription activator that may act as growth regulators in response to water deficit. Interacts with the core sequence 5'-CAATTATTA-3' of promoters in response to ABA and in an ABI1-dependent manner. Involved in the negative regulation of the ABA signaling pathway. {ECO:0000269|PubMed:10527431, ECO:0000269|PubMed:12065416}. |
| Regulation -- Description ? help
Back to Top |
| Source |
Description |
| UniProt | INDUCTION: By water deficit, by abscisic acid (ABA) and by salt stress. Self expression regulation. {ECO:0000269|PubMed:10527431, ECO:0000269|PubMed:12065416, ECO:0000269|PubMed:16055682}. |