 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | maker-scaffold00331-snap-gene-0.26-mRNA-1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Fagaceae; Castanea
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 279aa MW: 31400.1 Da PI: 4.6533 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 55.8 | 1e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT++Ed++l+ +++++G g+W++ ++ g+ R++k+c++rw +yl
maker-scaffold00331-snap-gene-0.26-mRNA-1 14 KGPWTPQEDQILISYIQKYGHGNWRALPKQAGLLRCGKSCRLRWTNYL 61
79******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 50.8 | 3.9e-16 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg+++ eE+e +++++++lG++ W++Ia++++ gRt++++k+ w+++l
maker-scaffold00331-snap-gene-0.26-mRNA-1 67 RGNFSIEEEETIIKLHEMLGNR-WSAIAARLP-GRTDNEIKNVWHTHL 112
89********************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.9E-26 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 17.581 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.76E-31 | 10 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 6.5E-15 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.3E-16 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.62E-12 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 25.336 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.7E-26 | 65 | 117 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.6E-13 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 5.6E-15 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.93E-10 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 279 aa Download sequence Send to blast |
MVRAPCCEKM GLKKGPWTPQ EDQILISYIQ KYGHGNWRAL PKQAGLLRCG KSCRLRWTNY 60 LRPDIKRGNF SIEEEETIIK LHEMLGNRWS AIAARLPGRT DNEIKNVWHT HLKKRLKQNQ 120 AYPETKLQPT NCDSNAPVQS ESSSLTAPSQ QGFESPAYTT MSPQTSSSDD ISSATDASMA 180 TMDQNNMTNN IKGEHIDIWE TFPTIDEEFW SDAPLAENIS MPSSLLPNND ELLQAQFPFN 240 STELVEPDYS DGSNIIGDGM DFWYNLFIGA GGSPELSEF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 1e-27 | 12 | 116 | 5 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 111 | 117 | LKKRLKQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00375 | DAP | Transfer from AT3G23250 | Download |
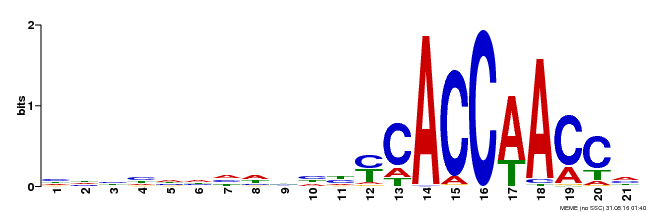
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_023879142.1 | 0.0 | transcription factor MYB14-like | ||||
| TrEMBL | A0A2N9GML5 | 1e-141 | A0A2N9GML5_FAGSY; Uncharacterized protein | ||||
| STRING | EOY06040 | 1e-120 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF31 | 34 | 817 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G23250.1 | 2e-77 | myb domain protein 15 | ||||



