 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | kfl00466_0010 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Klebsormidiophyceae; Klebsormidiales; Klebsormidiaceae; Klebsormidium
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 817aa MW: 84312.6 Da PI: 10.8206 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58.1 | 1.5e-18 | 68 | 119 | 6 | 56 |
S--HHHHHHHHHHHHH.SSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 6 tftkeqleeLeelFek.nrypsaeereeLAkklgLterqVkvWFqNrRakek 56
++t++q+++Le++F + + ++s+ ++ e+Ak +gL+ rqV +WFqNrRak++
kfl00466_0010 68 RLTTDQIHALEKAFDEaDGKLSHPQKVEIAKGIGLEARQVAIWFQNRRAKWR 119
689************99**********************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 95.4 | 5.7e-31 | 65 | 155 | 2 | 91 |
HD-ZIP_I/II 2 kkrrlskeqvklLEesFeeee.kLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreel 91
k rrl+++q+++LE+ F+e + kL++ +Kve+a+ +gl++rqva+WFqnrRA+++t q+E++y++Lk++ +l+++n+rL+ke ee r++l
kfl00466_0010 65 KIRRLTTDQIHALEKAFDEADgKLSHPQKVEIAKGIGLEARQVAIWFQNRRAKWRTEQVESQYDVLKAQNARLQADNDRLKKELEEARHAL 155
669**************99999**************************************************************9988776 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.11E-15 | 54 | 122 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 4.8E-17 | 59 | 126 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 15.92 | 60 | 121 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 8.2E-15 | 62 | 125 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 5.54E-14 | 67 | 122 | No hit | No description |
| Pfam | PF00046 | 6.7E-16 | 68 | 119 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 96 | 119 | IPR017970 | Homeobox, conserved site |
| CDD | cd14686 | 0.00365 | 114 | 149 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001558 | Biological Process | regulation of cell growth | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 817 aa Download sequence Send to blast |
MARPASASTP GASKRKREFG DAGKGKKSHK GKGKRSAEEA RGGPSAAEMR KMDSPAVQRA 60 KREIKIRRLT TDQIHALEKA FDEADGKLSH PQKVEIAKGI GLEARQVAIW FQNRRAKWRT 120 EQVESQYDVL KAQNARLQAD NDRLKKELEE ARHALVVARS GIAHPTPSRM EPVAGAGGAA 180 FFGSKDGESP GSVYEKLQNL LKSMKGKRSA FLAAAAAGES SDSGSGSSDE SSEEERQPIK 240 RAKRSALQKP RPALAAKPGP TPSRSSVAAG SGAKRDGTRR SPVGAPSKGS GADKGAGPSK 300 AVPKGRLTRE SSMGRPSTLQ RRGSKLEALQ LPVKMSPAVP LFADKDRAKA AVEAPTPRMG 360 RGKPSAATPG STPSMWLGPR QDPSPTTGPL APGEPQPAHP LGISGDGTVA LLDKVPPSAV 420 GRGFSHLLAG APESSPTDST VSPSAYLLSD GKEPSPTTGA LAGADAVPAK QSTPPKPSPL 480 REESGKEARL QLAPSASDWG EALRKAEQQR SAAAAAAAPA PARLDSPDLP TDMARDFVRA 540 LGAGEAARSP SPSVLPSVRE FPNSAAQDFL AKYRAMYGEI HRSSERTSEG ASKAADILAR 600 VARGGRRGGD TAGLAARLAA LDGGLSKGLD ALAVLHLGAA PGAGDNKGAA ERNGLLELSL 660 EPGSGLAPSR TAQLLAEAAA RVSAGSSLPS RLARGTAFLG SASAFEAFRG GGNLLDPYQD 720 PPFSSGLREE RALSPSLRDA ISQHRSGHTT QALLPGEGRE EAGAPGSSTG PLTPRLGPAA 780 PIPPWLTAVA SGLWGEGGSP GGRSSSPGWW TQIQSGH |
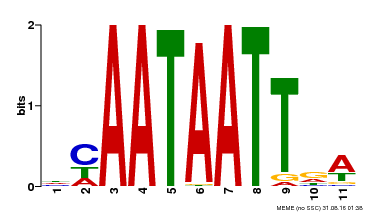
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00323 | DAP | Transfer from AT3G01470 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A1Y1IHB1 | 0.0 | A0A1Y1IHB1_KLENI; Homeobox protein | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP129 | 16 | 189 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G01470.1 | 1e-22 | homeobox 1 | ||||



