 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | kfl00264_0050 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Klebsormidiophyceae; Klebsormidiales; Klebsormidiaceae; Klebsormidium
|
||||||||
| Family | CPP | ||||||||
| Protein Properties | Length: 974aa MW: 100261 Da PI: 8.756 | ||||||||
| Description | CPP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCR | 46.1 | 1e-14 | 415 | 455 | 2 | 42 |
TCR 2 ekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNkeek 42
+kk+CnCk+s+Clk+YCeCfa+g++C+ eC+C++C+N+ e+
kfl00264_0050 415 KKKQCNCKNSRCLKLYCECFASGHYCGPECNCQGCCNNAEN 455
689**********************************9875 PP
| |||||||
| 2 | TCR | 53.4 | 5e-17 | 511 | 548 | 2 | 39 |
TCR 2 ekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNk 39
++kgC+Ckks ClkkYCeCf+ag+ C++ CkC dCkN
kfl00264_0050 511 HHKGCHCKKSGCLKKYCECFQAGVLCTDACKCIDCKNF 548
78***********************************5 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01114 | 2.6E-12 | 414 | 455 | IPR033467 | Tesmin/TSO1-like CXC domain |
| PROSITE profile | PS51634 | 35.141 | 415 | 550 | IPR005172 | CRC domain |
| Pfam | PF03638 | 2.0E-10 | 416 | 452 | IPR005172 | CRC domain |
| SMART | SM01114 | 3.3E-19 | 510 | 551 | IPR033467 | Tesmin/TSO1-like CXC domain |
| Pfam | PF03638 | 1.1E-13 | 513 | 548 | IPR005172 | CRC domain |
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 974 aa Download sequence Send to blast |
MEGGAVREQG GLVRPAAGAQ QQAGRLAQAG AAPASLPPRA GEGEWGMANN QAPRDDRTAH 60 TVPGSVRLAV QDGHGVGQIQ RPPHTVVLAA EEQPSRRQGQ AGQKHASTTA RPVGSPLPGA 120 LLPASAPPAG HSEAAGLGAS NGGMQAARNS FQGGARQQPD NATWEAPGGV GAAKPGAPQL 180 SSPGRRGDAP SAASHLAQPV RHPAPRFLGG SSQQGSAALR QPLPFGVHLM AGEAGGGRQH 240 QPAGSRQGLP SQGGLPSRGS AAPGSSPVTA WPSGPACPPS QLPSPRLIVR SMDPSHARAL 300 SPPGRPAPDG ALRKPVRHLD FGAQPYGLPA PPHQTAAHSW GPALPAASFL QGPSPHTAPL 360 PATSSSSPTW LLVRLELSLR ASPCVRCLLA ARPVPGASPY RGLRAAPDKP EPPPKKKQCN 420 CKNSRCLKLY CECFASGHYC GPECNCQGCC NNAENEVTRQ EAVEATLERN PNAFRPKIAP 480 SPDVKNEQEA AAPAAFLADP AATEAPVAGH HHKGCHCKKS GCLKKYCECF QAGVLCTDAC 540 KCIDCKNFDG SDERRLLLAG SIDPAPQGPP GYAAPSPPPF KKARLQDRAL QSPPQAKPAQ 600 GGSGAKSIPG SSYPAGRAPQ KPPTRSLLSG VVKADAVQEL CKLLAIVTQE VANNFEGPPT 660 PAPHGPLPTP VSSGLPVLQC DTDAESVFCL GSARCAAFCS CSPCAAAAFS RRGLTVAPCL 720 AGAAQHPGLL AAQQHRQEAA FAAAAARSDD GGRPGTSDDG AHTAVEERRA GGGSPAGQGP 780 GPGEAEEQSP EALLLSCDEE LPAEKEVQVA LPGGLYGGAD EGGGEGAAAR LYAAQEQTVL 840 EEFASCLKKI ISVGQKRLRQ YEETLSPMRP AYSGPPPLHY RPPPAPAGSA FPQRFPAGQA 900 PPASRQPGGA GRYVPQYHQQ APQQQQRGPA VGPAPYRPAY QQQQQQWQPA QAGRQARALP 960 RALRATGVVD RGIE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5fd3_A | 3e-30 | 416 | 558 | 11 | 131 | Protein lin-54 homolog |
| 5fd3_B | 3e-30 | 416 | 558 | 11 | 131 | Protein lin-54 homolog |
| Search in ModeBase | ||||||
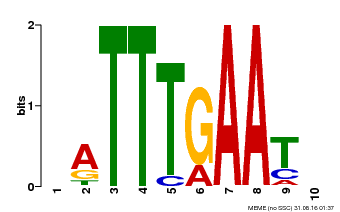
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00045 | PBM | Transfer from AT4G29000 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A0U9HSQ2 | 0.0 | A0A0U9HSQ2_KLENI; Uncharacterized protein | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP1469 | 17 | 45 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G29000.1 | 8e-38 | Tesmin/TSO1-like CXC domain-containing protein | ||||



