 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | kfl00079_0440 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Klebsormidiophyceae; Klebsormidiales; Klebsormidiaceae; Klebsormidium
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 402aa MW: 44099.2 Da PI: 6.5293 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 33.7 | 8.1e-11 | 111 | 146 | 3 | 38 |
XXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 3 elkrerrkqkNReAArrsRqRKkaeieeLeekvkeL 38
++k rr+++NReAAr+sR+RKka++++Le L
kfl00079_0440 111 PDKTLRRLAQNREAARKSRLRKKAYVQQLESSRIRL 146
68999************************9743333 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.20.5.170 | 7.2E-9 | 105 | 156 | No hit | No description |
| SMART | SM00338 | 5.6E-6 | 109 | 208 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 9.094 | 111 | 155 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 7.8E-8 | 111 | 152 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 5.63E-7 | 113 | 157 | No hit | No description |
| CDD | cd14708 | 4.29E-21 | 113 | 162 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 116 | 131 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF14144 | 1.8E-33 | 203 | 277 | IPR025422 | Transcription factor TGA like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009410 | Biological Process | response to xenobiotic stimulus | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 402 aa Download sequence Send to blast |
MASTLTPALA AGINSEVFSG KSQKTSDMSA PRDNNGAYDL ADLDGTPMNF GIAEDDDVGG 60 RGNGANGGAW NDTPMSPRSP SSDGGDMDEY KDQGSPSNKR PGDSDKKSEG PDKTLRRLAQ 120 NREAARKSRL RKKAYVQQLE SSRIRLQQLE QELQRARQQG VYHGDGAAGG GGTLPGLNSN 180 PNPGAAQFDL EYGRWVEEHS RQTSELRTAV NQRSTDDELR TLVDQCLAHY DELFKLKSQA 240 AKMDVFHLVS GMWKTPAERC FMWMGGFRPS EMLKIIIPQV EPLTEQQLMA ICNLQQSSQQ 300 AEDALSQGME SLQQSLADTL SAGNMSASSN VANYMGQMAM AMGKLGTLES FVRQADSLRQ 360 QSLQQMHRTL NIRQSAKGFL AMADYHARLR ALSSLWAARP RD |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-TGACG-3'. Recognizes ocs elements like the as-1 motif of the cauliflower mosaic virus 35S promoter. Binding to the as-1-like cis elements mediate auxin- and salicylic acid-inducible transcription. | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-TGACG-3'. Recognizes ocs elements like the as-1 motif of the cauliflower mosaic virus 35S promoter. Binding to the as-1-like cis elements mediate auxin- and salicylic acid-inducible transcription. Binds to the hexamer motif 5'-ACGTCA-3' of histone gene promoters. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00492 | DAP | Transfer from AT5G06950 | Download |
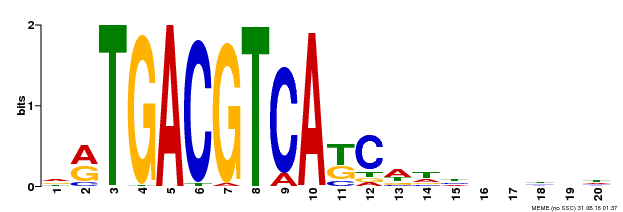
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002980439.2 | 1e-156 | transcription factor TGA2.3 isoform X2 | ||||
| Refseq | XP_024529114.1 | 1e-156 | transcription factor TGA2.3 | ||||
| Refseq | XP_024536795.1 | 1e-156 | transcription factor TGA2.3 isoform X1 | ||||
| Refseq | XP_024544912.1 | 1e-156 | transcription factor TGA2.3 isoform X2 | ||||
| Swissprot | O24160 | 1e-140 | TGA21_TOBAC; TGACG-sequence-specific DNA-binding protein TGA-2.1 | ||||
| Swissprot | P23923 | 1e-140 | HBP1B_WHEAT; Transcription factor HBP-1b(c38) | ||||
| TrEMBL | A0A1Y1HW44 | 0.0 | A0A1Y1HW44_KLENI; BZIP transcription factor | ||||
| STRING | EFJ22847 | 1e-159 | (Selaginella moellendorffii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP722 | 16 | 67 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06950.4 | 1e-111 | bZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



