 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | evm_27.model.AmTr_v1.0_scaffold00071.194 | ||||||||
| Common Name | AMTR_s00071p00185920 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; basal Magnoliophyta; Amborellales; Amborellaceae; Amborella
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 363aa MW: 42501.6 Da PI: 8.5585 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 17.1 | 1.6e-05 | 60 | 80 | 3 | 23 |
ETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 CpdCgksFsrksnLkrHirtH 23
C+dCg +F+ + +Lk+H + H
evm_27.model.AmTr_v1.0_scaffold00071.194 60 CEDCGACFRKPAYLKQHKQGH 80
*****************9877 PP
| |||||||
| 2 | zf-C2H2 | 17.9 | 8.4e-06 | 86 | 110 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++Cp C+ s++rk++L+rH+ +H
evm_27.model.AmTr_v1.0_scaffold00071.194 86 FTCPvdGCHSSYRRKDHLTRHLLKH 110
89*********************99 PP
| |||||||
| 3 | zf-C2H2 | 15 | 7.4e-05 | 115 | 139 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++Cp +C+ F+ + n krHi H
evm_27.model.AmTr_v1.0_scaffold00071.194 115 FECPvdNCNHKFTYQGNVKRHIEEH 139
89********************995 PP
| |||||||
| 4 | zf-C2H2 | 16.7 | 2.1e-05 | 155 | 179 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C+ Cgk F+ s L++H +H
evm_27.model.AmTr_v1.0_scaffold00071.194 155 HVCQepECGKAFKYESKLRKHEESH 179
689999***************9988 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 12 | 13 | 36 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.757 | 58 | 85 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.31 | 58 | 80 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 9.1E-7 | 60 | 84 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 60 | 80 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.65E-9 | 71 | 112 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-9 | 85 | 112 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0028 | 86 | 110 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.245 | 86 | 115 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 88 | 110 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 7.17E-6 | 96 | 145 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.0E-5 | 113 | 142 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.51 | 115 | 145 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.013 | 115 | 139 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 117 | 140 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.24E-5 | 152 | 180 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 7.9E-7 | 154 | 179 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.614 | 155 | 184 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.065 | 155 | 179 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 157 | 179 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 3.4 | 187 | 212 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 8.891 | 188 | 212 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 189 | 212 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 22 | 215 | 236 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 0.019 | 245 | 270 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.575 | 245 | 275 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.7E-4 | 246 | 271 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 247 | 270 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 8.42E-7 | 256 | 300 | No hit | No description |
| SMART | SM00355 | 5.2 | 276 | 300 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.162 | 276 | 300 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 278 | 300 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 363 aa Download sequence Send to blast |
MEREVIFKDV RRYYCDICGI CRSKKPLIRS HMLSEHKDEV NEVEKDEEEI LENKRLKLAC 60 EDCGACFRKP AYLKQHKQGH SLERPFTCPV DGCHSSYRRK DHLTRHLLKH EGKFFECPVD 120 NCNHKFTYQG NVKRHIEEHH SKESPSCSGQ DPLPHVCQEP ECGKAFKYES KLRKHEESHV 180 KLDSVEVICC DPECMKHFTN IEALREHLKS NHRYVLCEVC GSRQLKKNMK RHLRTHDNKV 240 SIERIKCPVR GCCHIFTTIS NLKKHEKAVH LELRPFTCRE PHCGRKFPYK HVRDNHEKSH 300 VYIHDDFVEL DEQFRSRPKG GRKPLFSGIE SLLRKRVAPL SRASSLDNAS EYLSSLLSED 360 GR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1tf6_A | 2e-16 | 54 | 214 | 11 | 162 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| 1tf6_D | 2e-16 | 54 | 214 | 11 | 162 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 224 | 230 | LKKNMKR |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
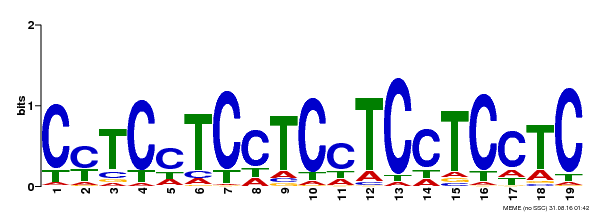
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020531805.1 | 0.0 | transcription factor IIIA isoform X2 | ||||
| Swissprot | Q84MZ4 | 1e-114 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | U5DF32 | 0.0 | U5DF32_AMBTC; Uncharacterized protein | ||||
| STRING | ERN20047 | 0.0 | (Amborella trichopoda) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP2981 | 16 | 27 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-110 | transcription factor IIIA | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | evm_27.model.AmTr_v1.0_scaffold00071.194 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



