 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | evm_27.model.AmTr_v1.0_scaffold00056.60 | ||||||||
| Common Name | AMTR_s00056p00085520 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; basal Magnoliophyta; Amborellales; Amborellaceae; Amborella
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 842aa MW: 92378.8 Da PI: 6.3481 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58.6 | 1.1e-18 | 17 | 75 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++++++ps +r++L +++ +++ +q+kvWFqNrR +ek+
evm_27.model.AmTr_v1.0_scaffold00056.60 17 KYVRYTPEQVEALERLYHECPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 75
5679*****************************************************97 PP
| |||||||
| 2 | START | 172.9 | 2e-54 | 161 | 369 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGC CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeq 71
+aee+++e+++ka+ ++ W +++ +++g++++ +++ s++++g a+ra+g+v +++ v+e+l+d++
evm_27.model.AmTr_v1.0_scaffold00056.60 161 IAEETLAEFLSKATGTAVEWIQMPGMKPGPDSIGIVAISHGCTGVAARACGLVGLEPT-KVSEILKDRPT 229
7899******************************************************.8999999999* PP
T-TT-SEEEEEEEECTT..EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--. CS
START 72 Wdetlakaetlevissg..galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe 138
W ++++++e+l ++s+g g+++l +++++a+++l++ Rdf+ +Ry+ l++g++v++++S++++q p+
evm_27.model.AmTr_v1.0_scaffold00056.60 230 WFRDCRSVEVLTALSTGngGTIELLYMQMYAPTTLASaRDFWLLRYTSVLEDGSLVVCERSLSNTQGGPS 299
******************************************************************9998 PP
...-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 139 ...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqce 205
+++vRae++pSg+li+p+++g+s ++ v+h+dl+ ++++++lr+l++s+++ ++k+++a+l+++++
evm_27.model.AmTr_v1.0_scaffold00056.60 300 mppVQQFVRAEMHPSGYLIRPCEGGGSIIHLVDHMDLEPWSVPEVLRPLYESSTVLAQKMTMAALRHLRQ 369
888899***********************************************************99876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.548 | 12 | 76 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.4E-16 | 14 | 80 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 6.68E-17 | 15 | 77 | No hit | No description |
| SuperFamily | SSF46689 | 2.91E-17 | 15 | 80 | IPR009057 | Homeodomain-like |
| Pfam | PF00046 | 2.4E-16 | 17 | 75 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-18 | 19 | 75 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 1.15E-6 | 69 | 108 | No hit | No description |
| PROSITE profile | PS50848 | 26.76 | 151 | 379 | IPR002913 | START domain |
| CDD | cd08875 | 1.16E-80 | 155 | 371 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 9.2E-26 | 160 | 366 | IPR023393 | START-like domain |
| SuperFamily | SSF55961 | 3.16E-40 | 160 | 372 | No hit | No description |
| SMART | SM00234 | 1.8E-41 | 160 | 370 | IPR002913 | START domain |
| Pfam | PF01852 | 4.6E-52 | 161 | 369 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 3.96E-5 | 397 | 494 | No hit | No description |
| SuperFamily | SSF55961 | 3.96E-5 | 521 | 601 | No hit | No description |
| Pfam | PF08670 | 1.1E-52 | 698 | 840 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 842 aa Download sequence Send to blast |
MAVTACKEGK PGMDPGKYVR YTPEQVEALE RLYHECPKPS SIRRQQLIRE CPILSNIEPK 60 QIKVWFQNRR CREKQRKEAS RLQAVNRKLT AMNKLLMEEN DRLQKQVSQL VYENGYFRQQ 120 TQNVAITTTD TSCESVVTSG QHHLTPQHPP RDASPAGLLS IAEETLAEFL SKATGTAVEW 180 IQMPGMKPGP DSIGIVAISH GCTGVAARAC GLVGLEPTKV SEILKDRPTW FRDCRSVEVL 240 TALSTGNGGT IELLYMQMYA PTTLASARDF WLLRYTSVLE DGSLVVCERS LSNTQGGPSM 300 PPVQQFVRAE MHPSGYLIRP CEGGGSIIHL VDHMDLEPWS VPEVLRPLYE SSTVLAQKMT 360 MAALRHLRQI AQEVSQNTVV GWGRQPAALR ALSQRLSKGF NEALNGFTDD GWSLLGNDGM 420 DDVTILVNSS PAKIMAANLT STNGFPTVCG AVLCAKASML LQNVPPALLI RFLREHRSEW 480 ADSNIDAYSA AALKSNPGTL PTSRMGGFGG QVILPLAHTV EHEEFLEVIK LENHGLVQDD 540 SIIPRDMFLL QLCSGVDENA VGACAELVFA PIDPSFADDA PLLPSGFRII PLDNGVDGSS 600 PNRTLDLASA LEVGPAGNRI SGEFAGGSGS LRSVLTIAFQ FSYENNHEQI RDNVATMARQ 660 YVRSVISSVQ RVAMALTPSR LNPHNGLRPP PGTPEALTLA RWICHSYRFH LGVELLRPNG 720 EGGESLLKML WHHADAIMCC SLKALPVFTF ANQAGLDMLE TTLVALQDIT LEKIFDENGR 780 KTLCADFAQI MQQGFAYLQG GLCVSSMGRP VSYERAVAWK VLNEEESTHC ICFMFMNWSF 840 V* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
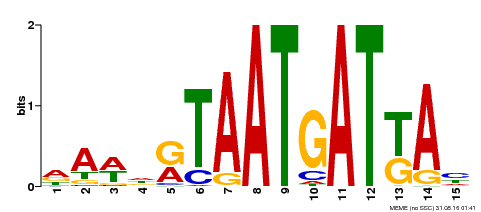
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006853643.2 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Refseq | XP_020528699.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | U5CYT1 | 0.0 | U5CYT1_AMBTC; Uncharacterized protein | ||||
| STRING | ERN15110 | 0.0 | (Amborella trichopoda) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP651 | 16 | 71 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | evm_27.model.AmTr_v1.0_scaffold00056.60 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



