 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | evm_27.model.AmTr_v1.0_scaffold00040.27 | ||||||||
| Common Name | AMTR_s00040p00057740 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; basal Magnoliophyta; Amborellales; Amborellaceae; Amborella
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 319aa MW: 35297.5 Da PI: 6.5489 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 45.5 | 1.6e-14 | 237 | 284 | 4 | 53 |
XCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 4 lkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelk 53
+r+rr++kNRe+A rsR+RK+a++ eLe+ v +Le+eN +L+ +++ +
evm_27.model.AmTr_v1.0_scaffold00040.27 237 QQRQRRMIKNRESAARSRERKQAYTVELETLVTQLEEENAQLQ--AQQVE 284
58****************************************8..33333 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.20.5.170 | 1.0E-13 | 229 | 295 | No hit | No description |
| SMART | SM00338 | 5.8E-11 | 234 | 295 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.795 | 236 | 281 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 7.0E-13 | 237 | 293 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14707 | 4.77E-19 | 238 | 292 | No hit | No description |
| SuperFamily | SSF57959 | 6.25E-10 | 239 | 281 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 241 | 256 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 319 aa Download sequence Send to blast |
MASKVMASSN STTNSDLNRQ ASIYSLTIAE LQNSNKGLGS MSMDDLLRNI WTAEESQAMA 60 AALAGNQPDS GQLAGNMSDS GQVMMNSGSF PHQNNLSLPK QVANKTVEEV WRELASSSAT 120 DQRGERMVYK EPTFGEMTLE DFLAKAGIVR EENQAQGVCE PSDWRRFGVK GDEMVSQQSM 180 FGDGVLGVQG QFQRLGGLEE GGLGSNNNNG LGLGLGHEGS SRGKRRAVHE PVDKVAQQRQ 240 RRMIKNRESA ARSRERKQAY TVELETLVTQ LEEENAQLQA QQVEENRKRR EELMKVLIPV 300 DEMPRPPRML RRTRSMHL* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00409 | DAP | Transfer from AT3G56850 | Download |
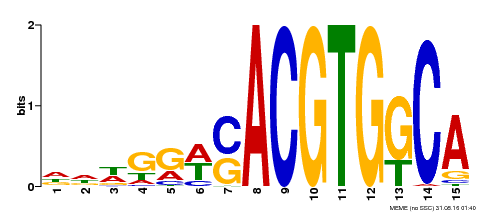
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020527287.1 | 0.0 | ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| TrEMBL | W1PY53 | 0.0 | W1PY53_AMBTC; Uncharacterized protein | ||||
| STRING | ERN12984 | 0.0 | (Amborella trichopoda) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP8662 | 7 | 15 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G41070.3 | 6e-25 | bZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | evm_27.model.AmTr_v1.0_scaffold00040.27 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



