 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | evm_27.model.AmTr_v1.0_scaffold00007.249 | ||||||||
| Common Name | AMTR_s00007p00238950 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; basal Magnoliophyta; Amborellales; Amborellaceae; Amborella
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 388aa MW: 42364.1 Da PI: 8.9993 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 41.9 | 2.5e-13 | 66 | 114 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s+ykGV + +grW A+I++ ++r++lg+f + eAa+a++ a++k++g
evm_27.model.AmTr_v1.0_scaffold00007.249 66 SRYKGVVPQP-NGRWGAQIYEK-----HQRVWLGTFNEEAEAARAYDTAALKFRG 114
78****9888.8*********3.....5**********99*************98 PP
| |||||||
| 2 | B3 | 99.3 | 2.4e-31 | 200 | 308 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT..........---..--SEEEEEETTS-EEEEEE..EEETTEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh..........ggkkeesktltledesgrsWevkliyrkksgry 59
f+k+ tpsdv+k++rlv+pk++ae+h gg +++ l++ed++g++W+++++y+++s++y
evm_27.model.AmTr_v1.0_scaffold00007.249 200 FEKAVTPSDVGKLNRLVIPKQHAERHfpldiassaaGGAGCKGVLLSFEDSTGKVWRFRYSYWNSSQSY 268
89***************************998888855555899************************* PP
EE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EEEEE-S CS
B3 60 vltkGWkeFvkangLkegDfvvFkldgrsefelvvkvfrk 99
vltkGW++Fvk++gLk+gD+v F++ e++l++ ++ +
evm_27.model.AmTr_v1.0_scaffold00007.249 269 VLTKGWSRFVKEKGLKAGDMVRFERGVGGEQQLYIDWRPR 308
************************7544777788888765 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 2.48E-17 | 66 | 122 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 1.1E-8 | 66 | 114 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 9.63E-25 | 66 | 121 | No hit | No description |
| SMART | SM00380 | 1.0E-28 | 67 | 128 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 20.56 | 67 | 122 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 4.3E-21 | 67 | 122 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 7.0E-39 | 196 | 309 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.99E-28 | 199 | 293 | No hit | No description |
| SuperFamily | SSF101936 | 3.79E-29 | 199 | 305 | IPR015300 | DNA-binding pseudobarrel domain |
| SMART | SM01019 | 1.2E-24 | 200 | 309 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 14.001 | 200 | 309 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 8.0E-29 | 200 | 308 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 388 aa Download sequence Send to blast |
MAESSANSTE QLAGIVIDSD DASKSPPPDD GASGLQRMGS GMSTVVDCEA GLGCIVEAES 60 RKLPSSRYKG VVPQPNGRWG AQIYEKHQRV WLGTFNEEAE AARAYDTAAL KFRGRDAVTN 120 FKSPRTSNHS HGNIGDDEEA EALFLATHSK GEIVDMLRKH TYHDELRQSK RYCRAMPVSR 180 ASATMPAEGR AVAAAREQLF EKAVTPSDVG KLNRLVIPKQ HAERHFPLDI ASSAAGGAGC 240 KGVLLSFEDS TGKVWRFRYS YWNSSQSYVL TKGWSRFVKE KGLKAGDMVR FERGVGGEQQ 300 LYIDWRPRPA AAPLLPPQFI VWQPAAAAPS PLAAVVHQPN VVRLFGVNLM TPPVAAPPQQ 360 VGFCLMKRGR EMEFFTCDAK KQCIGTL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 4e-52 | 197 | 311 | 11 | 120 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). Transcriptional repressor of flowering time on long day plants. Acts directly on FT expression by binding 5'-CAACA-3' and 5'-CACCTG-3 sequences (Probable). Functionally redundant with TEM1. {ECO:0000250, ECO:0000269|PubMed:18718758, ECO:0000305}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00025 | PBM | Transfer from AT1G68840 | Download |
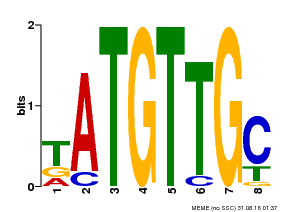
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006843763.2 | 0.0 | AP2/ERF and B3 domain-containing transcription repressor RAV2 | ||||
| Swissprot | P82280 | 1e-123 | RAV2_ARATH; AP2/ERF and B3 domain-containing transcription repressor RAV2 | ||||
| TrEMBL | W1PBW2 | 0.0 | W1PBW2_AMBTC; Uncharacterized protein | ||||
| STRING | ERN05438 | 0.0 | (Amborella trichopoda) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP365 | 16 | 108 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25560.1 | 1e-112 | RAV family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | evm_27.model.AmTr_v1.0_scaffold00007.249 |



