 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | evm.model.supercontig_4.49 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Caricaceae; Carica
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1536aa MW: 172410 Da PI: 8.9825 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 13 | 0.00031 | 1444 | 1466 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L++H r+H
evm.model.supercontig_4.49 1444 CPvkGCGKKFFSHKYLVQHRRVH 1466
9999*****************99 PP
| |||||||
| 2 | zf-C2H2 | 11.8 | 0.00075 | 1502 | 1528 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
evm.model.supercontig_4.49 1502 YVCAeaGCGQTFRFVSDFSRHKRKtgH 1528
89********************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 2.9E-16 | 23 | 64 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.853 | 24 | 65 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 2.1E-14 | 25 | 58 | IPR003349 | JmjN domain |
| SMART | SM00558 | 1.2E-49 | 200 | 369 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 33.752 | 203 | 369 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 4.94E-26 | 216 | 383 | No hit | No description |
| Pfam | PF02373 | 1.4E-36 | 233 | 352 | IPR003347 | JmjC domain |
| SMART | SM00355 | 23 | 1419 | 1441 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 0.0045 | 1442 | 1466 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.445 | 1442 | 1471 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.4E-5 | 1444 | 1465 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1444 | 1466 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.79E-9 | 1458 | 1500 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.7E-9 | 1466 | 1494 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0014 | 1472 | 1496 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.741 | 1472 | 1501 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1474 | 1496 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.26E-7 | 1490 | 1524 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.1E-8 | 1495 | 1525 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.91 | 1502 | 1528 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.448 | 1502 | 1533 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1504 | 1528 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1536 aa Download sequence Send to blast |
MAASTLSSDP TQEVFPWLKS LPLAPEFHPT FAEFQDPIAY IFKIEKEASK YGICKIVPPV 60 PPAPKKSAIT NLNRSLAARA ASAGAATSKP LPTFTTRQQQ IGFCPRKPRP VQKPVWQSGE 120 YYSFPEFEAK AKSFEKTYLK KCGKKGSLSP LEVETLYWKA TVDKPFSVEY ANDMPGSAFV 180 PVTGKKSVGV GGREAGEGVT VGETAWNMRG VSRAEGSLLR FMKEEIPGVT SPMVYIAMMF 240 SWFAWHVEDH DLHSLNYLHM GAGKTWYGVP RDAAVAFEEV VRVHGYGGEV NPLVTFATLG 300 EKTTVMSPEV FVSAGVPCCR LVQNAGEFVV TFPRAYHSGF SHGFNCGEAA NIATPEWLRV 360 AKDAAIRRAS INYPPMVSHF QLLYDLALAL CSRIPMNVSA KPRSSRLKDK KKGEGETLVK 420 ELFVQNVIQN NDLLHFLGKG SSAILLPEGS SDISVCSDLR VGSQLRAHPR MPLGACSYSD 480 VIKSSKGLDS GNDVLGSSDG MKPVKGTLAS QSLSKDTERE SSVQGDGLSD QRLFSCVTCG 540 ILSFACVAVV QPREAAARYL MSADCSFFND WTVSSGVTSE GFTLASRDTI ISEPRSRTRW 600 TEKSSQDGLF DVPVQSVDYQ IEMVNQANEG VSDTQMRGET SALGLLASTY GDSSDSDEDH 660 HATEARSPLL LRENSGDETS LRNIDCYEEP GYTRSNFKSR SDQTFDNSEF ETDNLTSPRS 720 NSLDSASKEL VMASNATSNC SPVSFVAEKT RFSRMIMPVE KADMPYAQKS DEDSSRLHVF 780 CLEHAVEVEK QLRPIGGVHI MLLCHPEYPR IEAEAKLVAE ELGIDYLWND IKFQVATRED 840 KERIQSALDS EEAIPGNGDW AVKLGINLFY SAVLSRSPMY TKQMPYNSVI YSAFGRSSPA 900 SSPSKSNSHG RRPAKQRKVV VGKWCGKVWM SNQVHPFLAQ RDLEEQEQER SFHARAIADE 960 NLENKPENIH KAQTTLETRK YSRKRKTATE IVPMKKIKPI ETEPVIPEGS LEDGSYKQHR 1020 RVSRSKQTKY VEKEDAVSCD SLEQNTRQWH RRSSRRKLAK TVEREDSLSD DPLEDNSLLK 1080 QRRILRGKRV EHFESDDAIS IGSLGDNSPL QQRRTLGSKQ TTFIEREDAG SDDDLVDDSH 1140 HHYQRISRGK ETQYVERDFT LSDDSMEDDS LQLDRRGLGN RQTTLTEMEG VVSCDSSEDH 1200 SQQQRSRTRR SKETKFVNKE DAGSYDSLED NSHQQPRKIP RGNQVKFIER EDTVSYDSLE 1260 DNCHPQGKRT QRSKKAKDIE REDGVSYDSL EDSSHQRRRR TLRSKKMKPK IIQKVKMETP 1320 QHMKKGKHLP AKQQTSRKKK QETPRQQNDK SGQNTRQFST YVEEQLEGGP STRLRKRILK 1380 SPKEIVVQVK EKKQTGKKKG KNASAGSSNA KVKDEEAEYL CDMEGCTMSF VSKQELVLHK 1440 RNICPVKGCG KKFFSHKYLV QHRRVHMDDR PLKCPWKGCK MTFKWAWART EHIRVHTGAR 1500 PYVCAEAGCG QTFRFVSDFS RHKRKTGHSA KKARG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 3e-73 | 17 | 390 | 8 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 3e-73 | 17 | 390 | 8 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 981 | 986 | SRKRKT |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
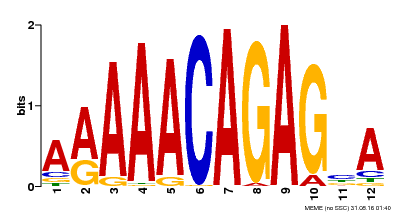
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | evm.model.supercontig_4.49 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021908879.1 | 0.0 | lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A061FYM0 | 0.0 | A0A061FYM0_THECC; Relative of early flowering 6, putative isoform 1 | ||||
| STRING | evm.model.supercontig_4.49 | 0.0 | (Carica papaya) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5259 | 28 | 43 | Representative plant | OGRP4401 | 12 | 17 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | evm.model.supercontig_4.49 |



