 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | evm.model.supercontig_2.88 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Caricaceae; Carica
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 560aa MW: 61007.6 Da PI: 9.3867 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 16.2 | 2.9e-05 | 71 | 93 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+ C+ C+k F r nL+ H r H
evm.model.supercontig_2.88 71 FICEVCNKGFQREQNLQLHRRGH 93
89*******************88 PP
| |||||||
| 2 | zf-C2H2 | 13.2 | 0.00027 | 147 | 169 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+kC++C+k++ +s+ k H +t+
evm.model.supercontig_2.88 147 WKCEKCSKRYAVQSDWKAHSKTC 169
58*****************9998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.160.60 | 6.2E-6 | 70 | 93 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 2.47E-7 | 70 | 93 | No hit | No description |
| PROSITE profile | PS50157 | 10.99 | 71 | 93 | IPR007087 | Zinc finger, C2H2 |
| Pfam | PF12171 | 2.7E-5 | 71 | 93 | IPR022755 | Zinc finger, double-stranded RNA binding |
| SMART | SM00355 | 0.0052 | 71 | 93 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 73 | 93 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 170 | 112 | 142 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 2.0E-5 | 135 | 168 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 2.47E-7 | 142 | 167 | No hit | No description |
| SMART | SM00355 | 130 | 147 | 167 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 560 aa Download sequence Send to blast |
MAASSSSVPL FGIREENQNQ MRQQQHSSTP APSAVQAPQP PKKKRNQPGT PNPDAEVIAL 60 SPKTLMATNR FICEVCNKGF QREQNLQLHR RGHNLPWKLK QKTTKEVKRK VYLCPDPTCV 120 HHDPSRALGD LTGIKKHYSR KHGEKKWKCE KCSKRYAVQS DWKAHSKTCG TREYRCDCGT 180 LFSRRDSFIT HRAFCDALAQ ESARHPPSIS TISSHLYGNN NNVAAAGLGL SQLGPQVPSL 240 QDQTTSQSTH APPTQFDHLL PSHSTFRHHH QQNPAASPAF FMQEPNQQDL LSNNKQFHGL 300 MQMAPGNNTT SLFNLGYNSS NDGGGKHSST TNLPSSGFLV PNHFNNENGT SGVDVVGGEG 360 STLFPNNLMT DQISSGLSSL FGTPIQGGSI NNMMPHMSAT ALLQKAAQMG STSSNNTASL 420 LRNKSNFASI FSENEANNLH ELMNSFSSGS PSILGGGYGE DHQNPYGEER QRHGPNTFGN 480 LDEQKLHQNM SANIGGSDRL TRDFLGVGQI VRSMSGGISQ REKQQQATMD ISSLESERNR 540 TGNAGPTTSQ SFGGGGNFQ* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5b3h_C | 8e-35 | 143 | 204 | 3 | 64 | Zinc finger protein JACKDAW |
| 5b3h_F | 8e-35 | 143 | 204 | 3 | 64 | Zinc finger protein JACKDAW |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor acting as a positive regulator of the starch synthase SS4. Controls chloroplast development and starch granule formation (PubMed:22898356). Binds DNA via its zinc fingers (PubMed:24821766). Recognizes and binds to SCL3 promoter sequence 5'-AGACAA-3' to promotes its expression when in complex with RGA (PubMed:24821766). {ECO:0000269|PubMed:22898356, ECO:0000269|PubMed:24821766}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
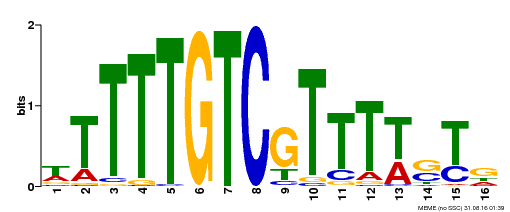
| Motif ID | Method | Source | Motif file |
| MP00255 | DAP | Transfer from AT2G02070 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | evm.model.supercontig_2.88 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AM455095 | 1e-120 | AM455095.1 Vitis vinifera contig VV78X055432.12, whole genome shotgun sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021909848.1 | 0.0 | protein indeterminate-domain 5, chloroplastic isoform X1 | ||||
| Swissprot | Q9ZUL3 | 1e-146 | IDD5_ARATH; Protein indeterminate-domain 5, chloroplastic | ||||
| TrEMBL | U5G1I0 | 0.0 | U5G1I0_POPTR; Uncharacterized protein | ||||
| STRING | evm.model.supercontig_2.88 | 0.0 | (Carica papaya) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4373 | 26 | 54 |
| Representative plant | OGRP95 | 16 | 242 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G02070.1 | 1e-143 | indeterminate(ID)-domain 5 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | evm.model.supercontig_2.88 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



