 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_17159_iso_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | WOX | ||||||||
| Protein Properties | Length: 278aa MW: 31226.9 Da PI: 5.6436 | ||||||||
| Description | WOX family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 67 | 2.5e-21 | 96 | 156 | 2 | 57 |
T--SS--HHHHHHHHHHHHH.SSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 2 rkRttftkeqleeLeelFek.nrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
r+R+t+t+ ql++Le+ F + n ps+++++e++++l +++e++V++WFqNrRa+ k+
cra_locus_17159_iso_1_len_1255_ver_3 96 RQRWTPTPVQLQILERIFDQgNGAPSKQKIKEITAELaqhgQISETNVYNWFQNRRARSKR 156
89*****************99**************************************97 PP
| |||||||
| 2 | Wus_type_Homeobox | 99.9 | 1.9e-32 | 96 | 158 | 3 | 65 |
Wus_type_Homeobox 3 rtRWtPtpeQikiLeelyksGlrtPnkeeiqritaeLeeyGkiedkNVfyWFQNrkaRerqkq 65
r+RWtPtp Q++iLe+++++G P+k++i++itaeL+++G+i+++NV++WFQNr+aR+++kq
cra_locus_17159_iso_1_len_1255_ver_3 96 RQRWTPTPVQLQILERIFDQGNGAPSKQKIKEITAELAQHGQISETNVYNWFQNRRARSKRKQ 158
89***********************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 14.916 | 92 | 157 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 6.2E-18 | 92 | 163 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 3.2E-14 | 94 | 161 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.84E-18 | 94 | 161 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 3.95E-14 | 96 | 158 | No hit | No description |
| Pfam | PF00046 | 3.7E-19 | 96 | 156 | IPR001356 | Homeobox domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 278 aa Download sequence Send to blast |
MEWDKQQQPQ PPPPSAEEQL NGGTANNNNG GMFVKVMTDE QMEVLRKQIA VYATICEQLV 60 DLHKSLTSHH DLAGVRLGNL YCDPLVTSAG HKITGRQRWT PTPVQLQILE RIFDQGNGAP 120 SKQKIKEITA ELAQHGQISE TNVYNWFQNR RARSKRKQQA AASNNVESEV ETEVESPNEK 180 KTKPEDLQSQ HMPTSRADDL CFQNTDVSSV MHSMDPRNSK AEPMFPSEGS SKPIGSFGQM 240 PFYGSMLPNT RMEQLLGKME VPESYSPYLQ AEDYNMTG |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 149 | 157 | RRARSKRKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor which may be involved in developmental processes. {ECO:0000250}. | |||||
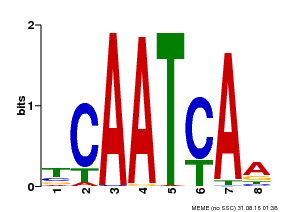
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00669 | PBM | Transfer from GRMZM2G038252 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027076418.1 | 1e-162 | WUSCHEL-related homeobox 8-like isoform X1 | ||||
| Swissprot | O81788 | 4e-88 | WOX13_ARATH; WUSCHEL-related homeobox 13 | ||||
| TrEMBL | A0A068TZT5 | 1e-161 | A0A068TZT5_COFCA; Uncharacterized protein | ||||
| STRING | PGSC0003DMT400003980 | 1e-154 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA5675 | 24 | 37 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



