|
Plant Transcription
Factor Database
|
Transcription Factor Information
|
Basic
Information? help
Back to Top |
| TF ID |
Zpz_sc01238.1.g00070.1.sm.mk |
| Organism |
|
| Taxonomic ID |
|
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
| Family |
C2H2 |
| Protein Properties |
Length: 438aa MW: 47803.9 Da PI: 8.1127 |
| Description |
C2H2 family protein |
| Gene Model |
| Gene Model ID |
Type |
Source |
Coding Sequence |
| Zpz_sc01238.1.g00070.1.sm.mk | genome | ZGD | View Nucleic Acid |
|
| Signature Domain? help Back to Top |
 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | zf-C2H2 | 17.4 | 1.2e-05 | 84 | 106 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C++C+k F r nL+ H r H
Zpz_sc01238.1.g00070.1.sm.mk 84 FVCEICNKGFQRDQNLQLHRRGH 106
89*******************88 PP
|
| 2 | zf-C2H2 | 12.2 | 0.00057 | 206 | 228 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C+ Cgk++ +s+ k H + +
Zpz_sc01238.1.g00070.1.sm.mk 206 WRCERCGKRYAVHSDWKAHVKGC 228
58*****************9866 PP
|
| 3 | zf-C2H2 | 10.7 | 0.0017 | 236 | 254 | 5 | 23 |
TTTEEESSHHHHHHHHHHT CS
zf-C2H2 5 dCgksFsrksnLkrHirtH 23
dCg Fsrk++L++H +
Zpz_sc01238.1.g00070.1.sm.mk 236 DCGILFSRKDSLMTHRAFC 254
8**************8765 PP
|
| Sequence ? help Back to Top |
| Protein Sequence Length: 438 aa
Download sequence Send
to blast |
MLSDLSSDQE ATGSSSHGGD MASYVLSSPL FLAPAAATAL LPQPPPPEQP RAGAKRKRSQ 60
PGNPDPDAEV VALSPRTLVA TNRFVCEICN KGFQRDQNLQ LHRRGHNLPW KLRQRTAPSG 120
SGGGGRQQGE AAQPPRKRVY VCPEPTCVHH DPTRSIEKAC CSIETIKNEI LHASWYVIQL 180
TLMYRALGDL TGIKKHFSRK HGEKRWRCER CGKRYAVHSD WKAHVKGCGT KEYRCDCGIL 240
FSRKDSLMTH RAFCDALAEE SARLLAAAHH NSSTITSING GGSDNNNLLI TSNSTPLFLP 300
FSNPPSAADH NPNNPLFFLP QQEAPLLQQI HHSTYLDLPV VDATVSTAAS VTGDSVADTI 360
SFGLIPEGSV AALHAAGDVG QRRLTRDFLG VDHSEVEELQ LPLCATVPRV ASCATDLTRQ 420
YFVGERLPPV NNTWSNNF
|
| Functional Description ? help
Back to Top |
| Source |
Description |
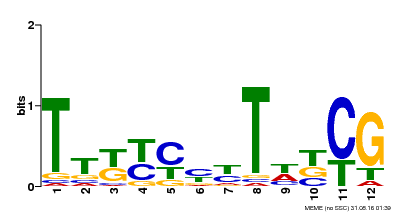
| UniProt | Transcription activator that acts as a flowering master switch in both long and short days, independently of the circadian clock (PubMed:18790997, PubMed:18725639, PubMed:18774969, PubMed:19304997, PubMed:24280027). Promotes flowering upstream of HD1 by up-regulating FTL1, FTL4, FTL5, FTL6, EHD1, HD3A and RFT1 (PubMed:18790997, PubMed:18725639, PubMed:18774969). Seems to repress FTL11 expression (PubMed:18725639). May recognize the consensus motif 5'-TTTGTCGTAAT-3' in target gene promoters (PubMed:18725639). {ECO:0000269|PubMed:18725639, ECO:0000269|PubMed:18774969, ECO:0000269|PubMed:18790997, ECO:0000269|PubMed:24280027, ECO:0000303|PubMed:19304997}. |
| Regulation -- Description ? help
Back to Top |
| Source |
Description |
| UniProt | INDUCTION: Induced by FKF1 (PubMed:25850808). Down-regulated by DLF1 (PubMed:25036785). During the basic vegetative growth phase (BVP, photoperiod-insensitive phase), suppressed via phytochrome-mediated light signals involving the phytochromobilin synthase HY2 (PubMed:25573482). {ECO:0000269|PubMed:25036785, ECO:0000269|PubMed:25573482, ECO:0000269|PubMed:25850808}. |
| Annotation --
Nucleotide ? help
Back to Top |
| Source |
Hit ID |
E-value |
Description |
| GenBank | AF058757 | 6e-76 | AF058757.1 Zea mays zinc finger protein ID1 (id1) mRNA, complete cds. |
| GenBank | BT053802 | 6e-76 | BT053802.1 Zea mays full-length cDNA clone ZM_BFb0184E01 mRNA, complete cds. |
| Publications
? help Back to Top |
- Rice Chromosome 10 Sequencing Consortium
In-depth view of structure, activity, and evolution of rice chromosome 10.
Science, 2003. 300(5625): p. 1566-9
[PMID:12791992] - Park SJ, et al.
Rice Indeterminate 1 (OsId1) is necessary for the expression of Ehd1 (Early heading date 1) regardless of photoperiod.
Plant J., 2008. 56(6): p. 1018-29
[PMID:18774969] - Song Y,Gao Z,Luan W
Interaction between temperature and photoperiod in regulation of flowering time in rice.
Sci China Life Sci, 2012. 55(3): p. 241-9
[PMID:22527521] - Hu S, et al.
A point mutation in the zinc finger motif of RID1/EHD2/OsID1 protein leads to outstanding yield-related traits in japonica rice variety Wuyunjing 7.
Rice (N Y), 2013. 6(1): p. 24
[PMID:24280027] - Cai Y, et al.
Dlf1, a WRKY transcription factor, is involved in the control of flowering time and plant height in rice.
PLoS ONE, 2014. 9(7): p. e102529
[PMID:25036785] - Yoshitake Y, et al.
The effects of phytochrome-mediated light signals on the developmental acquisition of photoperiod sensitivity in rice.
Sci Rep, 2015. 5: p. 7709
[PMID:25573482] - Han SH,Yoo SC,Lee BD,An G,Paek NC
Rice FLAVIN-BINDING, KELCH REPEAT, F-BOX 1 (OsFKF1) promotes flowering independent of photoperiod.
Plant Cell Environ., 2015. 38(12): p. 2527-40
[PMID:25850808] - Jin J, et al.
MORF-RELATED GENE702, a Reader Protein of Trimethylated Histone H3 Lysine 4 and Histone H3 Lysine 36, Is Involved in Brassinosteroid-Regulated Growth and Flowering Time Control in Rice.
Plant Physiol., 2015. 168(4): p. 1275-85
[PMID:25855537] - Brambilla V,Gomez-Ariza J,Cerise M,Fornara F
The Importance of Being on Time: Regulatory Networks Controlling Photoperiodic Flowering in Cereals.
Front Plant Sci, 2017. 8: p. 665
[PMID:28491078]
|