 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zpz_sc00289.1.g00260.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 587aa MW: 63366.6 Da PI: 11.0864 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 101.4 | 8.3e-32 | 364 | 456 | 2 | 103 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEE CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFk 82
F+ k+y++++d++++ ++ w +n+f+vld+ f++ +Lp+yFkh+nfaSFvRQLn+YgF+kv+ ++ weF+
Zpz_sc00289.1.g00260.1.am.mk 364 FVAKTYHMVSDPATDAVVCWGVASNTFLVLDPAAFSDYLLPSYFKHRNFASFVRQLNTYGFRKVDADR---------WEFA 435
9********************999*****************************************999.........**** PP
ESXXXXXXXXXXXXXXXXXXX CS
HSF_DNA-bind 83 hksFkkgkkellekikrkkse 103
h+sF +g+++ll i rk+++
Zpz_sc00289.1.g00260.1.am.mk 436 HESFLRGQTQLLPLIVRKRKK 456
****************99875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 1.4E-34 | 355 | 447 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM00415 | 5.8E-48 | 360 | 453 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| SuperFamily | SSF46785 | 1.09E-29 | 361 | 453 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| PRINTS | PR00056 | 2.0E-16 | 364 | 387 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Pfam | PF00447 | 1.2E-27 | 364 | 453 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 2.0E-16 | 402 | 414 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 403 | 427 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 2.0E-16 | 415 | 427 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0006970 | Biological Process | response to osmotic stress | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 587 aa Download sequence Send to blast |
MKGKRGAAGK LRCTLGGARF CSTTGAASRR SLGPGSIRHE VQGLHVCHDV STRGMLAPLP 60 RRKAKAGSLS GKSSRCKQHP SGSTPLARRL EKRCDADADA DSPKRATPRG RVRNVAARGC 120 RGLSPPPGSL PLGHLTARLL APPHEERRGW LPGRGALGWL GAGAFDGHGH DRRSGDKRCA 180 ATRTESAQNP SRPQRHITPQ IRPLHLLPAS GGAPPSPRHM ARPCAEADKV GTRRSALRPG 240 PSEIYIGGNA FRCRGQKYLL CGRVGVAPHR RRSRINPTPT PTAHTQSGAK QPKRKSQRLS 300 YPVLPPSCAR ALSRVRDTGS VCSSYLAVAS RALVAMGSEC YVQLQAKQQQ PPPAQAGAGA 360 IAPFVAKTYH MVSDPATDAV VCWGVASNTF LVLDPAAFSD YLLPSYFKHR NFASFVRQLN 420 TYGFRKVDAD RWEFAHESFL RGQTQLLPLI VRKRKKPGCR ELCDEGEEVR GTMKAVQRLR 480 EEQRGMDEEL QAMDRRLQAA ESRPGQMMTF LAKLADEPGA VLRAMLAKKE ELAEAGRKGS 540 TAPLEAPGKR RRIGGGEDCG AVAGGDAAEQ GMGAVQFPFS VLGQVFY |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2ldu_A | 9e-27 | 356 | 453 | 12 | 120 | Heat shock factor protein 1 |
| 5d5u_B | 9e-27 | 353 | 453 | 18 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 9e-27 | 353 | 453 | 18 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 9e-27 | 353 | 453 | 18 | 129 | Heat shock factor protein 1 |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 451 | 455 | RKRKK |
| 2 | 548 | 552 | KRRRI |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00379 | DAP | Transfer from AT3G24520 | Download |
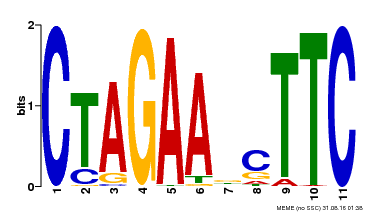
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Zpz_sc00289.1.g00260.1.am.mk |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK066316 | 1e-152 | AK066316.1 Oryza sativa Japonica Group cDNA clone:J013066G03, full insert sequence. | |||
| GenBank | AY344493 | 1e-152 | AY344493.1 Oryza sativa (japonica cultivar-group) heat shock factor RHSF11 mRNA, complete cds. | |||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP11298 | 31 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G24520.1 | 2e-55 | heat shock transcription factor C1 | ||||



