 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma75g00700.1 | ||||||||
| Common Name | ZOSMA_75G00700 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1017aa MW: 115317 Da PI: 6.7089 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 177.6 | 1.6e-55 | 22 | 138 | 2 | 118 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfq 95
++ ++rwl++ ei++iL nf+++ ++ e+++rp+sgs++L++rk++ryfrkDGy+w+kkkdgktv+E+he+LK g+++vl+cyYah+een +fq
Zosma75g00700.1 22 IDAQSRWLRPAEICEILRNFQNFCIAPEPPNRPPSGSIFLFDRKILRYFRKDGYNWRKKKDGKTVKEAHERLKAGSIDVLHCYYAHGEENDKFQ 115
56799***************************************************************************************** PP
CG-1 96 rrcywlLeeelekivlvhylevk 118
rr+yw+Lee++ +ivlvhy+evk
Zosma75g00700.1 116 RRSYWMLEEKYMHIVLVHYREVK 138
********************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 81.093 | 17 | 143 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 9.1E-76 | 20 | 138 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 7.5E-50 | 24 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 3.7E-6 | 432 | 520 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 4.0E-7 | 435 | 520 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 1.87E-17 | 435 | 520 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 1.87E-17 | 618 | 734 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 3.5E-17 | 620 | 736 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 3.50E-17 | 620 | 732 | No hit | No description |
| PROSITE profile | PS50297 | 18.033 | 627 | 734 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 8.7E-8 | 645 | 735 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.36 | 673 | 702 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 19 | 712 | 741 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 9.03E-8 | 834 | 893 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.25 | 842 | 864 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.018 | 844 | 863 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.206 | 844 | 872 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.025 | 865 | 887 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.194 | 866 | 890 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 1.2E-4 | 869 | 887 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1017 aa Download sequence Send to blast |
MSGNQLCAST TAQQLDVEQI LIDAQSRWLR PAEICEILRN FQNFCIAPEP PNRPPSGSIF 60 LFDRKILRYF RKDGYNWRKK KDGKTVKEAH ERLKAGSIDV LHCYYAHGEE NDKFQRRSYW 120 MLEEKYMHIV LVHYREVKGN KTSHTKGVVP HSVYDHSPVA SNSLTIRNQI TSQTTDANYP 180 DSVQISGYED SKMDNFQASS RHYPIPEMSQ FANLMDASLP KTCFPVFQNT QGDSEERGTT 240 VFDETKYEIT HNETKSQMGL TSLDEVLEYC NVEHQTHFPS TFHNNLESIP TRRKSVVSNH 300 EVSTGKLSSG IKDVANTYGQ ENLQFPGPDS FVMVSKKDAD NEINDDHCSI LNQPSTINDV 360 QGDKSLKKYD SFSRWMSKEL GQMDNSNMQS SSETYWDVAD SEGVVEDFSS SRQDDLYPYP 420 YVLSPSLSQD QYFSIIDFSP NWAYSDMETK VIITGTFIMS KIEVERCKWS CMFGEVEVPA 480 DALSDGILRC YAPPHKVGRV PFYVTCSNRL ACSEVREFEY LVSHAQYMEN SETYAGSKNE 540 MLLHIRLEEL LSLGLGTYES LELKMCHANY EKSLIRNQIS SLLLKSDERY ALAKKTNEIK 600 FSSRTSKDEL MQKLLKEKLH AWLLCKVVED GKGPNILDKE GQGVIHLAAG LGYDWAIKPI 660 VLAGVNINFR DARGWSALHW AAYFGRERTV AMLISLDTAP GALTDPTPEF PNGRTPADLA 720 SSNGHKGIAG YLAESSLTTH LSVLTLRDSL QTESIDIEDT TERHANKFDE RTVQTGSSLE 780 DSLSAVRSAT LAAASIQQVF RVQSFHRKKL VESSDTKCRM VGEHALSLVA IKTNKTRHSN 840 EPVHAAAIRI QNSFRGWKGK KEFTVIRQHI VKIQAHVRGH QVRKHFRKVV WSVGIVEKVI 900 LRWRRKGSGL RGFRSNALTK ISDIQIPSKD DGGDEDYDFL REGRKQTEQR MNRALSRVKS 960 MVQYPEARDQ YRRLLTGVEE LKDDAPGGTL EKETDDLMMF DVTELLGEED VHMSPA* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
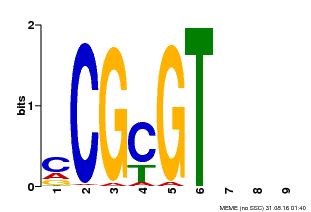
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_029119343.1 | 0.0 | calmodulin-binding transcription activator 3 isoform X2 | ||||
| TrEMBL | A0A0K9NPW7 | 0.0 | A0A0K9NPW7_ZOSMR; Calmodulin-binding transcription activator 3 | ||||
| STRING | XP_008795547.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma75g00700.1 |



