 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma293g00070.1 | ||||||||
| Common Name | ZOSMA_293G00070 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 366aa MW: 40027.7 Da PI: 7.9382 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 37.7 | 3.7e-12 | 302 | 346 | 7 | 55 |
HHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 7 erErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+Er RR ri +++ +L+el+P+ +k + a++L Av+YIk+Lq
Zosma293g00070.1 302 VAERVRRTRISERMRKLQELVPNM----DKQTNTADMLDFAVDYIKDLQ 346
78*********************7....688*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50888 | 14.799 | 295 | 345 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 2.36E-15 | 298 | 357 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 7.70E-12 | 299 | 346 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 3.4E-14 | 300 | 356 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 2.2E-12 | 301 | 351 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 4.8E-9 | 302 | 346 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 366 aa Download sequence Send to blast |
MYGSPSMSKD LNLPFPLSPY KQAKEDPDVR RQKQQPHVSS SGLLRYRTAP GNFLGEVCDE 60 TVATESMFAR FMGPDVRDEE KHHQSSTNHE VAITGEFPSS DPPHMFFHSP LQHHQPQMSS 120 SISPVGTGPL DREAATGGGA GNLVRHSSSP VGFLSHSNSE NGFAGIRSMS GFGGSSEGRT 180 SSGRLKGQIS FSSQHISELG SESMDRSSPG EGGSNRCYIP GYPIVSLGDS VPISDEFGTV 240 GRKRGIRDPS ESQNHVPTLM HHFSLPKTAT EVAATERIMQ FQDFVPCKIR AKRGCATHPR 300 SVAERVRRTR ISERMRKLQE LVPNMDKQTN TADMLDFAVD YIKDLQGEAK MLSENRAKCT 360 CSRSD* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 26 | 33 | PDVRRQKQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00646 | PBM | Transfer from LOC_Os08g39630 | Download |
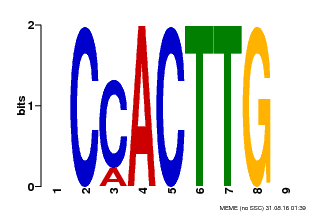
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020691604.1 | 1e-108 | transcription factor bHLH130 | ||||
| TrEMBL | A0A0K9PC85 | 0.0 | A0A0K9PC85_ZOSMR; Basic helix-loop-helix (BHLH) DNA-binding superfamily | ||||
| STRING | XP_008776552.1 | 1e-102 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1884 | 38 | 103 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G42280.1 | 2e-60 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma293g00070.1 |



