 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma174g00410.1 | ||||||||
| Common Name | ZOSMA_174G00410 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | BES1 | ||||||||
| Protein Properties | Length: 298aa MW: 32276.1 Da PI: 8.2319 | ||||||||
| Description | BES1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF822 | 187.3 | 6.3e-58 | 3 | 124 | 2 | 130 |
DUF822 2 gsgrkptwkErEnnkrRERrRRaiaakiyaGLRaqGnyklpkraDnneVlkALcreAGwvvedDGttyrkgskpleeaeaagssasaspessl 94
++gr+ twkErEnnkrRERrRRaiaaki++GLRa+Gn+klpk++DnneVlkALc +AGw+ve+DGttyrkgskp ++e++g+ as ++ +
Zosma174g00410.1 3 SGGRMATWKERENNKRRERRRRAIAAKIFSGLRAYGNFKLPKHCDNNEVLKALCVDAGWTVEEDGTTYRKGSKPA-ATETVGNGASFMISPLY 94
689************************************************************************.77777777666655444 PP
DUF822 95 qsslkssalaspvesysaspksssfpspssldsisl 130
+ a++sp++sy+asp+sssfpsp+++++ s+
Zosma174g00410.1 95 S------AFPSPAPSYHASPSSSSFPSPTRVENSSA 124
4......99*********************987654 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05687 | 2.4E-56 | 3 | 125 | IPR008540 | BES1/BZR1 plant transcription factor, N-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009742 | Biological Process | brassinosteroid mediated signaling pathway | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048316 | Biological Process | seed development | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 298 aa Download sequence Send to blast |
MPSGGRMATW KERENNKRRE RRRRAIAAKI FSGLRAYGNF KLPKHCDNNE VLKALCVDAG 60 WTVEEDGTTY RKGSKPAATE TVGNGASFMI SPLYSAFPSP APSYHASPSS SSFPSPTRVE 120 NSSAPPTVPN TEPYFLPHFR NLPPLRISNS APVTPPLSSP TCRPAKMLKP DWATYANSFH 180 HPIFAASEPS SPTRRNGHPA PIPECDESDA STVDSDRWGV NFQMTAPPSP TFNLVNPTAT 240 SLPETIAMAV VSERGGIVGS EFQFECGSVK AWAGERIHEF GVDDLELTLG NGKHNHG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zd4_A | 8e-23 | 6 | 83 | 372 | 449 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_B | 8e-23 | 6 | 83 | 372 | 449 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_C | 8e-23 | 6 | 83 | 372 | 449 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_D | 8e-23 | 6 | 83 | 372 | 449 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00237 | DAP | Transfer from AT1G75080 | Download |
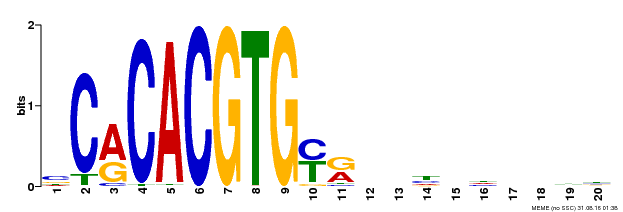
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009411027.1 | 2e-95 | PREDICTED: protein BZR1 homolog 1 | ||||
| Swissprot | Q94A43 | 1e-82 | BEH2_ARATH; BES1/BZR1 homolog protein 2 | ||||
| TrEMBL | A0A0K9PS01 | 0.0 | A0A0K9PS01_ZOSMR; BES1/BZR1-like protein | ||||
| STRING | XP_009349751.1 | 2e-94 | (Pyrus x bretschneideri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5474 | 34 | 53 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G75080.2 | 1e-67 | BES1 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma174g00410.1 |



