 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc01961.1.g00030.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 740aa MW: 79738.4 Da PI: 9.2409 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 129.6 | 1.2e-40 | 82 | 158 | 2 | 78 |
-SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 2 CqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
C v+gC+adls+++eyhrrhkvCe+hsk+pvv+v+g+e rfCqqCsrfh+l efDe+krsCr+rL++hn+rrrk+q+
Zmw_sc01961.1.g00030.1.am.mk 82 CAVDGCDADLSKCREYHRRHKVCEAHSKTPVVVVAGREMRFCQQCSRFHSLVEFDEAKRSCRKRLDGHNRRRRKPQP 158
**************************************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 9.5E-35 | 74 | 143 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.129 | 79 | 156 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.57E-37 | 80 | 161 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 8.8E-31 | 82 | 155 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 740 aa Download sequence Send to blast |
MWPGAMLWCL FSEHKGSMPL LKGEHGPGAP LGPSPLESLQ PSTSRKLCIC CASSISLASS 60 LSLRARPRTG PGGANGPQCP SCAVDGCDAD LSKCREYHRR HKVCEAHSKT PVVVVAGREM 120 RFCQQCSRFH SLVEFDEAKR SCRKRLDGHN RRRRKPQPDD MNARSFMTSQ QGPRSSSFVT 180 PRPEPSWSGI LKSEDSSYYA HQVLCNRPHF AGSTSTYSKE GRRFPFLQDG DQLSFGTGGA 240 ASLEAATACQ SLLKTVAAPE SSSSSKIFSP DGLTPVLDSD CALSLLSSPA NSSSVDVSRM 300 VQPTEHIPMA QPLVPSMQQF GSSPSWFSQA ASSGVVVSTA GFTCPSMDSE QLNPVLVPSS 360 DGHEMNYHGI FHVGGEGSDD GTSSSLPFSS TTSVEVHTKP DAAVSNLEQM IQFRPEAAVK 420 HGTIPLCSPD AALLSSLLDP QRVCLCSLAT HGRQGLGRWL VPPQCTAVGT RQGRLARKPL 480 RPDCIPRGAG GRQGTKFCLA RPGAGRGRAD KASEHGKMSV LGVQASKVEW HVTSRGAEQP 540 PRATGRPPDL GIVDRHVVKL MMGSCRVRAP YDVAPHPAIP PPTARVRARY PADAAGGPSN 600 RWFRTADDGV RVHNLGLAPL AEGWREVSVL YSLVPLVRTE KPSRLTLTGE LRPVCSNCGN 660 QSLASSRAEP ERQPRRPVAG LEEAGNDAAH GKRKGGVAQK YDSTRFHPEK YTLYALFFIS 720 LPRPVESSLV RRRDLQTLVV |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 2e-27 | 70 | 155 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 138 | 155 | KRSCRKRLDGHNRRRRKP |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
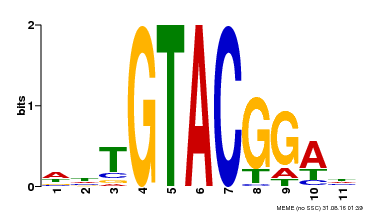
| Motif ID | Method | Source | Motif file |
| MP00555 | DAP | Transfer from AT5G50570 | Download |
| |||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1755 | 37 | 95 |



