 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc01752.1.g00150.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 180aa MW: 19011.6 Da PI: 10.436 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 41.4 | 3.5e-13 | 10 | 46 | 1 | 42 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeA 42
++y+GVr ++ +gr++AeIrdp + r+r++lg+f++ eA
Zmw_sc01752.1.g00150.1.am.mk 10 PRYRGVRKRP-WGRFAAEIRDPAK---RARVWLGTFDS-AEA 46
68********.**********843...6**********.555 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 1.1E-8 | 10 | 46 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 2.01E-21 | 10 | 67 | No hit | No description |
| SMART | SM00380 | 3.2E-22 | 11 | 71 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 6.5E-22 | 11 | 65 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 2.09E-14 | 11 | 66 | IPR016177 | DNA-binding domain |
| PROSITE profile | PS51032 | 16.199 | 11 | 65 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 5.1E-8 | 12 | 23 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 5.1E-8 | 34 | 50 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 180 aa Download sequence Send to blast |
MCDTAAVAAP RYRGVRKRPW GRFAAEIRDP AKRARVWLGT FDSAEAXXXX AARNLRGPLA 60 RTNFPVVPRL PLTSSRPRGL VVAPAQATAP TCSSSSTVES SSGPRVGPKT AAAPRKRVVV 120 VLPRPAAPDV GCHSDCASST SVMDDGDDAS TVRSRVPFDL NLAPSMDGDL DAGTDLRLCL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gcc_A | 3e-15 | 12 | 66 | 3 | 61 | ETHYLENE RESPONSIVE ELEMENT BINDING FACTOR 1 |
| 2gcc_A | 3e-15 | 12 | 66 | 6 | 64 | ATERF1 |
| 3gcc_A | 3e-15 | 12 | 66 | 6 | 64 | ATERF1 |
| Search in ModeBase | ||||||
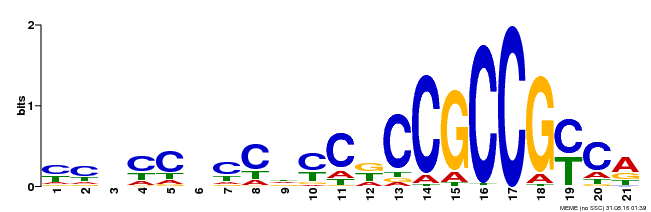
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00366 | DAP | Transfer from AT3G20310 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025822790.1 | 2e-49 | ethylene-responsive transcription factor 7-like | ||||
| TrEMBL | A0A0A8Y8Q6 | 5e-65 | A0A0A8Y8Q6_ARUDO; Uncharacterized protein | ||||
| STRING | Pavir.Ga00187.1.p | 1e-54 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5123 | 27 | 58 |



