 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc01401.1.g00130.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 773aa MW: 82118.8 Da PI: 10.3266 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 128.4 | 2.8e-40 | 296 | 372 | 1 | 77 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S-- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkq 77
+CqvegC++ l+ akeyhrrhkvCe+hskap v+v g eqrfCqqCsrfh+lsefD++krsCrrrLa+hnerrrk++
Zmw_sc01401.1.g00130.1.am.mk 296 RCQVEGCHVLLAGAKEYHRRHKVCEAHSKAPRVVVLGAEQRFCQQCSRFHALSEFDDAKRSCRRRLAGHNERRRKSS 372
6**************************************************************************87 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 2.3E-33 | 290 | 358 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.505 | 294 | 371 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.15E-37 | 295 | 374 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 1.4E-31 | 297 | 370 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0010229 | Biological Process | inflorescence development | ||||
| GO:0010321 | Biological Process | regulation of vegetative phase change | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 773 aa Download sequence Send to blast |
MYSVRSTVHG RRVHVLYSTQ HQLQRQRQRT AGNCAGSSQN RGLRTAYPLL PNAQHGDPIS 60 PKARTYSPFP SQSDMGRAVA DTDDWTGPYA LRQARTLRPC TPGLLPRAGF PASGSSVPCV 120 MTGGSESRQT VALLLSGGLA RLPACLPACF LCGEPRRVNL PPYLQQQHQT PPLFRLCLPR 180 RRQEAMEGNG GGSGTATPSW DLGMHWAPAG SSGYPQPFAP QLGGGPSRDQ QQHQQEQTWL 240 NLGKQPCCWA GAGGSQQAPQ LSSWGHVHGN GGGTGGGSGA AAAAEGKRKE KAAAPRCQVE 300 GCHVLLAGAK EYHRRHKVCE AHSKAPRVVV LGAEQRFCQQ CSRFHALSEF DDAKRSCRRR 360 LAGHNERRRK SSASETMART VGHPHGLMPF GHGFFPSSPA GALSLLSSAR GGGAPWLIPA 420 TADISARSSA ALDELIAENR RALLAWQFFS DRSPAGRLVP SSSTGRVLPA GEVPAGWHTA 480 QIQDHPGGSG GRADDRYSFS EAPAGAGHTT LDLMQTPDTS AGAPFLQPVS ERAGNARTPR 540 KDGSDAGCTS VDAWASAEGA PDRVHRCPPR PARAESQNRS GSSSVVSLNH PLQSRRRRGR 600 RPSQTNSGAR LITVPTCRSE LWDGGASYVS PRSAMMAIMA SPMRGSAIGF GKCGPYVVCK 660 ESHKRETTYA AMASPSAFRF DPAAEVTDES RQNTTTRRQA WAAAARGYGG WSYGGATWYG 720 SPYGAGGDAI FTSDQGGRKP SSDPLPLLWA AMGQFSSRAA EAAARVPVEP GRG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 3e-29 | 297 | 370 | 11 | 84 | squamosa promoter binding protein-like 4 |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
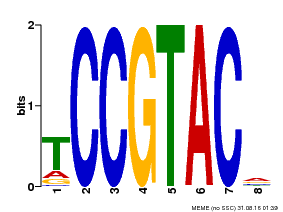
| Motif ID | Method | Source | Motif file |
| MP00290 | DAP | Transfer from AT2G33810 | Download |
| |||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP10303 | 34 | 39 |



