 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zjn_sc00051.1.g00100.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 610aa MW: 65452.4 Da PI: 5.1546 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 56.4 | 7.5e-18 | 47 | 96 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
++ GVr+++ +grWvAeI+d + ++ r +lg+f taeeAa+a+++a+ l+g
Zjn_sc00051.1.g00100.1.am.mk 47 KFVGVRQRP-SGRWVAEIKDTT---QKIRMWLGTFETAEEAARAYDEAACLLRG 96
799******.7*********32...35**********************99998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 1.24E-18 | 47 | 104 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 1.8E-12 | 47 | 96 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 3.96E-30 | 47 | 103 | No hit | No description |
| PROSITE profile | PS51032 | 19.836 | 47 | 104 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.2E-33 | 47 | 110 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 2.5E-27 | 47 | 104 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.6E-7 | 48 | 59 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.6E-7 | 70 | 86 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000302 | Biological Process | response to reactive oxygen species | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0035865 | Biological Process | cellular response to potassium ion | ||||
| GO:0048528 | Biological Process | post-embryonic root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 610 aa Download sequence Send to blast |
MELQFQKPQS HAQQQQCQYQ AVSVVAKETA GKLSRARSKC GGGGGGKFVG VRQRPSGRWV 60 AEIKDTTQKI RMWLGTFETA EEAARAYDEA ACLLRGANTR TNFAAAGGAG SAPPVSPLAS 120 RIRALLHHKK LKKTVARQPD VAFSPPAAAA YRHGGGTGSV AATSTSASTV TTTTSSVSPS 180 SSPSSTTIKF AMTSGGIRTP ILPVPSHHTF TEESTCYRPY LLTTGSDEFQ HQYEQSWTLN 240 TSSLPPTDGR DNACSRMPEL EKISPEKDGP ASPPHGEDRV QGGQDFFDAG NDASDSFLAG 300 ESLSPAVIAA DQLAIAMATA GVSSAAERQR PSCKEEVVQL SRLLGDSPSI LALQRFSSAM 360 DRRVAPSMDA GQDVWVWEVL LPDDHTSSST TCLLDTTNLS DDQETKEPMM ITPPSSDTDV 420 RDEPVDEFKD IGVVLDESTE PSEEAALPEA TDLAAASDGD EGEKPFQSAD TKKPDEVADP 480 SAASDGKEEE KQFQSRDTKE SVDDGEFEGE EDDAIKNKKG NGQERPECAV FSVGKLKVNG 540 IGALCSFGVA AATVCVFLIG GKLHHHHHRH QQQRRQKIQL QFDDKTRVLH SPTPCDLQRI 600 QQVVQQTSKG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wx9_A | 4e-13 | 42 | 112 | 9 | 80 | Ethylene-responsive transcription factor ERF096 |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00521 | DAP | Transfer from AT5G19790 | Download |
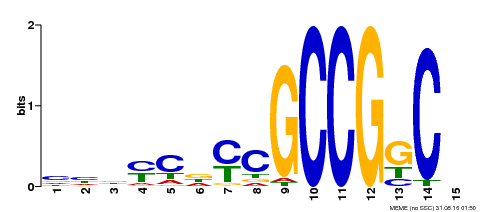
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP004273 | 3e-57 | AP004273.2 Oryza sativa Japonica Group genomic DNA, chromosome 7, PAC clone:P0431A02. | |||
| GenBank | AP014963 | 3e-57 | AP014963.1 Oryza sativa Japonica Group DNA, chromosome 7, cultivar: Nipponbare, complete sequence. | |||
| GenBank | CP012615 | 3e-57 | CP012615.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 7 sequence. | |||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3900 | 32 | 73 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G19790.1 | 2e-22 | related to AP2 11 | ||||



