 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zjn_sc00029.1.g03690.1.sm.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 454aa MW: 49145.6 Da PI: 7.4427 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 41.6 | 2.3e-13 | 312 | 358 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++Pk + K++Ka+ L A+ YI++L+
Zjn_sc00029.1.g03690.1.sm.mk 312 NHVEAERQRREKLNQRFYALRAVVPKiS------KMDKASLLSDAIAYIQELE 358
799***********************66......****************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 1.5E-39 | 42 | 212 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 17.047 | 308 | 357 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.38E-14 | 311 | 362 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 3.0E-16 | 311 | 361 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 1.96E-16 | 311 | 363 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 7.0E-11 | 312 | 358 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 8.1E-16 | 314 | 363 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009611 | Biological Process | response to wounding | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 454 aa Download sequence Send to blast |
MSWSETDAAL FTAVLGRDAA HHLATTPPHV DGPATAALAP ELQDRLHDLV ERGGAWTYGI 60 FWQESRIVLG WGDGHCRDGE AAHGDTTEAQ SAARKSALLR LHALYGGGDD DDGADYALRL 120 DRVTGAEMYF LASMYFSFPE GVGGPGRALA SRRHAWAAVD PRGPVPGWFV RASLAQAAGL 180 RTVVFLPCNG GVLELGSVVP VRETPEALRA IEAALAVTPA APEESMRIFG KDLSRRGQLP 240 AAAQANWAPP QLGGQTSPDD RKKDAVEQKP RDPPPKRVDF TKLEPNGQAV VEERKPRKRG 300 RKPANGREEP LNHVEAERQR REKLNQRFYA LRAVVPKISK MDKASLLSDA IAYIQELEDR 360 LRGGAKSPPA VEVKAMQDDV VLRVTTPLYA HPASRVFHAI RDARLSVVAS ELAVADDAVT 420 HTLMLRSPGP ERLTAEAVLA AMSRGTEGRD TPSP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 1e-24 | 306 | 361 | 2 | 57 | Transcription factor MYC2 |
| 5gnj_B | 1e-24 | 306 | 361 | 2 | 57 | Transcription factor MYC2 |
| 5gnj_E | 1e-24 | 306 | 361 | 2 | 57 | Transcription factor MYC2 |
| 5gnj_F | 1e-24 | 306 | 361 | 2 | 57 | Transcription factor MYC2 |
| 5gnj_G | 1e-24 | 306 | 361 | 2 | 57 | Transcription factor MYC2 |
| 5gnj_I | 1e-24 | 306 | 361 | 2 | 57 | Transcription factor MYC2 |
| 5gnj_M | 1e-24 | 306 | 361 | 2 | 57 | Transcription factor MYC2 |
| 5gnj_N | 1e-24 | 306 | 361 | 2 | 57 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 293 | 301 | RKPRKRGRK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00101 | PBM | Transfer from AT2G46510 | Download |
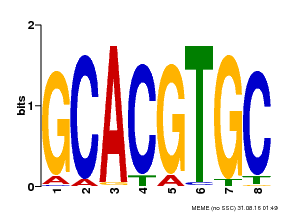
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KJ727698 | 1e-134 | KJ727698.1 Zea mays clone pUT5637 bHLH transcription factor (bHLH57) gene, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015628703.1 | 0.0 | transcription factor bHLH13 | ||||
| TrEMBL | A2ZX10 | 0.0 | A2ZX10_ORYSJ; Uncharacterized protein | ||||
| STRING | ONIVA01G32020.1 | 0.0 | (Oryza nivara) | ||||
| STRING | ORGLA01G0232100.1 | 0.0 | (Oryza glaberrima) | ||||
| STRING | OBART01G28080.1 | 0.0 | (Oryza barthii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP7385 | 29 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16430.1 | 5e-63 | bHLH family protein | ||||



