 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zjn_sc00019.1.g00440.1.sm.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | M-type_MADS | ||||||||
| Protein Properties | Length: 270aa MW: 29340.7 Da PI: 8.6321 | ||||||||
| Description | M-type_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 56.3 | 4e-18 | 11 | 50 | 3 | 42 |
--SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-T CS
SRF-TF 3 ienksnrqvtfskRrngilKKAeELSvLCdaevaviifss 42
i n+s r tf kRr+g++KKA+EL +LCd++++v+ +++
Zjn_sc00019.1.g00440.1.sm.mk 11 IVNDSTRRATFKKRRKGLMKKASELATLCDVDACVVAYGE 50
7799**********************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00432 | 2.0E-21 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS50066 | 17.731 | 1 | 50 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00266 | 7.92E-33 | 2 | 85 | No hit | No description |
| PRINTS | PR00404 | 3.3E-10 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 2.35E-24 | 3 | 95 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 8.7E-18 | 11 | 50 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 3.3E-10 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 3.3E-10 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0043078 | Cellular Component | polar nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 270 aa Download sequence Send to blast |
MARKKVTLQW IVNDSTRRAT FKKRRKGLMK KASELATLCD VDACVVAYGE GESQPEVWPS 60 VAEAARILAR FKAMPELDQC KKMMDMEGFL KQRIDKLREQ LHKAQRENRE RETTLLLHDA 120 IVGRRPGLVG LSIEELTSLG SMTDERLQNV NKALENLQAQ GGQGLPATAL QLQLSQAASM 180 TLPPLVTYPA PGVGQRDVQA PPNMQCWMID VPKPGGELGA MLYGGGFGGG STQQGFDGAG 240 SFTAGGHMTQ LGNMGAGFQW PDPGQSFPPM |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 2 | 24 | RKKVTLQWIVNDSTRRATFKKRR |
| 2 | 21 | 25 | KKRRK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00553 | DAP | Transfer from AT5G48670 | Download |
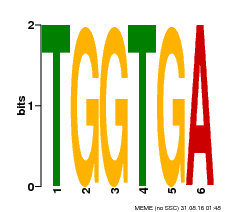
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KJ727716 | 1e-149 | KJ727716.1 Zea mays clone pUT5655 MADS transcription factor (MADS64) gene, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008646559.1 | 1e-109 | agamous-like MADS-box protein AGL80 | ||||
| TrEMBL | A0A1E5ULN7 | 1e-120 | A0A1E5ULN7_9POAL; Agamous-like MADS-box protein AGL80 | ||||
| STRING | GRMZM5G853066_P01 | 1e-108 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP363 | 36 | 202 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G48670.1 | 2e-40 | AGAMOUS-like 80 | ||||



