 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zjn_sc00013.1.g08240.1.sm.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 349aa MW: 37731.1 Da PI: 5.7926 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 59.6 | 7.5e-19 | 11 | 61 | 2 | 56 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkr..krfslgkfgtaeeAakaaiaarkklege 56
+++GVr+++ +grW+AeIrdp + +r +lg+f taeeAa a++ a +l+g+
Zjn_sc00013.1.g08240.1.sm.mk 11 HFRGVRQRP-WGRWAAEIRDP-----HhrRRLWLGTFNTAEEAAVAYDSASIRLHGA 61
69*******.**********7.....447*************************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 9.81E-22 | 11 | 69 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 1.3E-29 | 11 | 69 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 23.354 | 11 | 68 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 3.15E-32 | 11 | 70 | No hit | No description |
| Pfam | PF00847 | 2.4E-12 | 11 | 60 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.4E-32 | 11 | 74 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.2E-10 | 12 | 23 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.2E-10 | 34 | 50 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 349 aa Download sequence Send to blast |
MEALGPRPDV HFRGVRQRPW GRWAAEIRDP HHRRRLWLGT FNTAEEAAVA YDSASIRLHG 60 AGATTNFPTA RYLPPLEPAE PIVSLSPAPG TVITLPPVPV NPTVPLQLEK DVGSCEDQGE 120 VGATTLVEVP VHQQIVSLTP APGTLITLPP VPVNPTVPQQ VMDEVGSCED QEELGATSLV 180 EVPGRQQIVS LTHASWLPQT LPHTLPPVPV NPTVPQQVMK EVGSCEDQEE VGATSLVEVP 240 VHQQIVSLPG LLITLLPVPG NPTVPLLVMK EVGSCEDQEE VGATSLVEMP VHQQMPLRET 300 IAGHALRAQQ ARWAHASHRY HTWAMPTSCA ALCPGGRTTF HKHVIVLHN |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 3e-21 | 11 | 68 | 5 | 63 | ATERF1 |
| 3gcc_A | 3e-21 | 11 | 68 | 5 | 63 | ATERF1 |
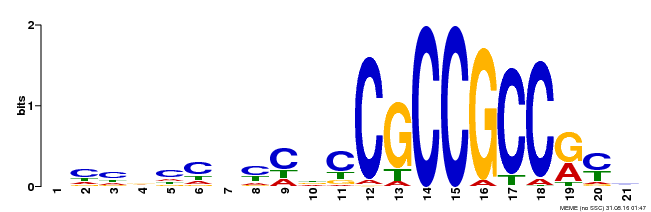
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00124 | DAP | Transfer from AT1G03800 | Download |
| |||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP11573 | 28 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G03800.1 | 7e-27 | ERF domain protein 10 | ||||



