 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zjn_sc00008.1.g08940.1.sm.mkhc | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 374aa MW: 40477.8 Da PI: 9.4191 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 179.7 | 7.8e-56 | 27 | 157 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpkkvkaeekewyfFskrdkkyatgkrknratk 79
lppGfrFhPtdeel+++yLk+k+++ ++++ ++i+evd++k+ePwdLp+k+k +ekewyfFs rd+ky+tg r+nrat+
Zjn_sc00008.1.g08940.1.sm.mkhc 27 LPPGFRFHPTDEELITYYLKQKLADGSFTA-RAIAEVDLNKCEPWDLPEKAKLGEKEWYFFSLRDRKYPTGVRTNRATN 104
79*************************999.89***************99999************************** PP
NAM 80 sgyWkatgkdkevlskkgel.....vglkktLvfykgrapkgektdWvmheyrl 128
+gyWk+tgkdke+++ ++ vg+kktLvfykgrap+gekt+Wvmheyr+
Zjn_sc00008.1.g08940.1.sm.mkhc 105 AGYWKTTGKDKEIYTG-QQPatpelVGMKKTLVFYKGRAPRGEKTNWVMHEYRV 157
**************95.33345556***************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 7.32E-64 | 24 | 178 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 60.539 | 27 | 178 | IPR003441 | NAC domain |
| Pfam | PF02365 | 4.5E-29 | 28 | 157 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 374 aa Download sequence Send to blast |
MSAEGSGGTA AGAGGGGGGG SKKEESLPPG FRFHPTDEEL ITYYLKQKLA DGSFTARAIA 60 EVDLNKCEPW DLPEKAKLGE KEWYFFSLRD RKYPTGVRTN RATNAGYWKT TGKDKEIYTG 120 QQPATPELVG MKKTLVFYKG RAPRGEKTNW VMHEYRVHSK SISKSTKDEW VVCRVFAKSA 180 GVKKYPSNNS HSRSHHHLHH PYTLDMVPPL VPTLLQHDPF ARHHHHPYMT PADMAELARL 240 ARGTPGLHPH IQPHPGGFIN PTAPQFAIPG LNLNLGASPA MPSPPPPQPA LHAAMTTSMA 300 AMGHHTAPSA GGPGGHVMGS EQQQQMAPGL GGCVMAPGAG PYGGFAGTDG VRYQNLDVEQ 360 LVERYWPGTA GYQV |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 6e-56 | 26 | 184 | 16 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 6e-56 | 26 | 184 | 16 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 6e-56 | 26 | 184 | 16 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 6e-56 | 26 | 184 | 16 | 171 | NO APICAL MERISTEM PROTEIN |
| 3swm_A | 8e-56 | 26 | 184 | 19 | 174 | NAC domain-containing protein 19 |
| 3swm_B | 8e-56 | 26 | 184 | 19 | 174 | NAC domain-containing protein 19 |
| 3swm_C | 8e-56 | 26 | 184 | 19 | 174 | NAC domain-containing protein 19 |
| 3swm_D | 8e-56 | 26 | 184 | 19 | 174 | NAC domain-containing protein 19 |
| 3swp_A | 8e-56 | 26 | 184 | 19 | 174 | NAC domain-containing protein 19 |
| 3swp_B | 8e-56 | 26 | 184 | 19 | 174 | NAC domain-containing protein 19 |
| 3swp_C | 8e-56 | 26 | 184 | 19 | 174 | NAC domain-containing protein 19 |
| 3swp_D | 8e-56 | 26 | 184 | 19 | 174 | NAC domain-containing protein 19 |
| 4dul_A | 6e-56 | 26 | 184 | 16 | 171 | NAC domain-containing protein 19 |
| 4dul_B | 6e-56 | 26 | 184 | 16 | 171 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the promoter regions of genes involved in chlorophyll catabolic processes, such as NYC1, SGR1, SGR2 and PAO. {ECO:0000269|PubMed:27021284}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00277 | DAP | Transfer from AT2G24430 | Download |
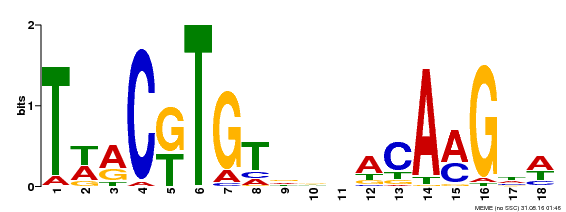
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Repressed by the microRNA miR164. {ECO:0000269|PubMed:15294871, ECO:0000269|PubMed:17098808}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025803276.1 | 0.0 | NAC domain-containing protein 92-like | ||||
| Swissprot | Q9FLJ2 | 1e-80 | NC100_ARATH; NAC domain-containing protein 100 | ||||
| TrEMBL | A0A3L6QN81 | 0.0 | A0A3L6QN81_PANMI; Uncharacterized protein | ||||
| STRING | LPERR09G11330.1 | 0.0 | (Leersia perrieri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP9361 | 38 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G24430.2 | 7e-89 | NAC domain containing protein 38 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



