 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_016492244.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 481aa MW: 53261.3 Da PI: 6.706 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 54.1 | 2.8e-17 | 301 | 347 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn ErrRRdriN+++ L+ellP++ +K +Ka++L +A+eY+ksLq
XP_016492244.1 301 VHNLSERRRRDRINEKMKALQELLPHS-----TKTDKASMLDEAIEYLKSLQ 347
6*************************9.....9******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50888 | 18.476 | 297 | 346 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 3.01E-20 | 300 | 374 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 2.79E-18 | 300 | 351 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 4.2E-20 | 301 | 355 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.2E-14 | 301 | 347 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 5.0E-18 | 303 | 352 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0009693 | Biological Process | ethylene biosynthetic process | ||||
| GO:0010244 | Biological Process | response to low fluence blue light stimulus by blue low-fluence system | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0010928 | Biological Process | regulation of auxin mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 481 aa Download sequence Send to blast |
MNPCLPEWNI EAELSVPHQK KPIGFDHELV ELLWRNGEVV LHSQTHKKQP GYDPNECRQF 60 NKHDQQTIRD VVSCGNHTNL IQDDETISWL NCPIDESFEK EFCSPFLSEI STNPIEVDKS 120 IRQSEDNKAF KFDTLEINNV FPHSHHSGFD PNPMPPPRFH NTGSAKQHNV KEGSVMTVGS 180 SHCGSNQVAI DADTSRFSSS ANIGLSAAMI TDYTGKVSPQ SDTMDRDTFE PANTSSSGGS 240 GSSYARTCNL SAATNSQSHK RKSRDGEEPE CQSKAGELES AGGNKSAQKS GTARRSRAAE 300 VHNLSERRRR DRINEKMKAL QELLPHSTKT DKASMLDEAI EYLKSLQMQL QMMWMGSGMA 360 SMMFPGVQHY MSRMGMGMGP PTVPSIHNAM HLARLPLVDS AIPLNPAAPN QAAMCQNSML 420 NSVNYQRHLQ NPNFQDQYAS YMGFHPLQGA SQQMNIFGLG SQTAQQSQQL SHPTNSNAPN 480 T |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 305 | 310 | ERRRRD |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00082 | ChIP-seq | Transfer from AT3G59060 | Download |
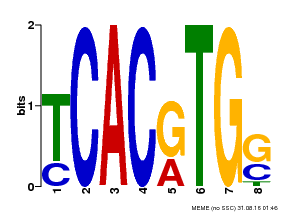
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT013582 | 0.0 | BT013582.1 Lycopersicon esculentum clone 132330F, mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009602219.1 | 0.0 | PREDICTED: transcription factor PIF4 | ||||
| Refseq | XP_016492244.1 | 0.0 | PREDICTED: transcription factor PIF4-like | ||||
| Refseq | XP_018626747.1 | 0.0 | PREDICTED: transcription factor PIF4 | ||||
| Refseq | XP_018626748.1 | 0.0 | PREDICTED: transcription factor PIF4 | ||||
| Refseq | XP_018626749.1 | 0.0 | PREDICTED: transcription factor PIF4 | ||||
| TrEMBL | A0A1S4BTN6 | 0.0 | A0A1S4BTN6_TOBAC; transcription factor PIF4-like | ||||
| STRING | XP_009602219.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA10465 | 22 | 26 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43010.2 | 2e-51 | phytochrome interacting factor 4 | ||||



