 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_016478076.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1103aa MW: 123690 Da PI: 6.0312 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 182.4 | 5.2e-57 | 20 | 136 | 2 | 118 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqr 96
l+ ++rwl++ ei++iL+n++k++++ e+++rp+sgsl+L++rk++ryfrkDG+sw+kkkdgktv+E+he+LK g+++vl+cyYah+een++fqr
XP_016478076.1 20 LEAQHRWLRPAEICEILKNYQKFRIAPEPPNRPPSGSLFLFDRKVLRYFRKDGHSWRKKKDGKTVKEAHERLKAGSIDVLHCYYAHGEENENFQR 114
5679******************************************************************************************* PP
CG-1 97 rcywlLeeelekivlvhylevk 118
r+yw+Leee+++ivlvhy+evk
XP_016478076.1 115 RSYWMLEEEMSHIVLVHYREVK 136
*******************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 83.558 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.9E-79 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 3.7E-50 | 21 | 134 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 5.3E-4 | 517 | 600 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 9.57E-16 | 518 | 602 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 4.6E-4 | 518 | 594 | IPR002909 | IPT domain |
| PROSITE profile | PS50297 | 17.582 | 700 | 822 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 4.76E-12 | 700 | 809 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 1.8E-17 | 701 | 814 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.48E-16 | 705 | 812 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 6.8E-8 | 722 | 813 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 1.9 | 750 | 780 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 8.576 | 750 | 783 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 120 | 790 | 819 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 3.45E-7 | 919 | 976 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 50 | 925 | 947 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.346 | 927 | 955 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.18 | 928 | 946 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0042 | 948 | 970 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.285 | 949 | 973 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 5.0E-4 | 951 | 970 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1103 aa Download sequence Send to blast |
MADSRRYGLN AQLDIDQILL EAQHRWLRPA EICEILKNYQ KFRIAPEPPN RPPSGSLFLF 60 DRKVLRYFRK DGHSWRKKKD GKTVKEAHER LKAGSIDVLH CYYAHGEENE NFQRRSYWML 120 EEEMSHIVLV HYREVKGNRT NFSRTREPQE ATPRFQETDE DVHSSEVDSS ASTKFYPNGY 180 QVNSQVTDAT SLSSAQASEY EDAESAYNQH PTSGFHSFLD AQPSMMQKAG ESLPVPYHPI 240 PFSTDDHQVQ FAGSSDMDFF SSAPGNKSRN TANTYIPSRN LDFPSWETIS VNNPAAYQSY 300 HFQPSSQSGA NNMTHEQGST TMGQVFLNDF KKQGQNRIDS LGDWQTSEGD AAFISKWSMD 360 QKLNPNLASD HTIRSSAAYN VELHNSLEAS HILPSHQDKH PMQNELPSQL SDANVGGSLN 420 AELDHNLSIG VRTDHSSLKQ PLLDGVLREG LKKLDSFDRW MSKELEDVSE PHMQSNSSSY 480 WDNVGDDDGV DNSTIASQVQ LDTYMLSPSL SQDQFFSIID FSPSWAFAGS EIKVLITGKF 540 LKSQPEVEKW ACMFGELEVP AEVIADGVLR CHTPNQKVGR VPFYITCSNR LACSEVREFE 600 FRVSESQDVD VANSCSSSES LLHMRFGKLL SLESTVSLSS PPRSEDDVSN VCSKINSLLK 660 EDDNEWEEML NLTYENNFMA EKVKDQLLQK LLKEKLRVWL LQKVAEGGKG PNVLDEGGQG 720 VLHFAAALGY DWAIPPTIAA GVSVNFRDVN GWTALHWAAS YGRERTVGFL IISLGAAPGA 780 LTDPTPKHPS GRTPADLASS NGHKGIAGYL AESSLSSHLS SLELKEMKQG ETVQPFGEAV 840 QTVSERSATP AWDGDWPHGV SLKDSLAAVR NATQAAARIH QVFRVQSFQR KQLKEHGGSE 900 FGLSDEHALS LLALKTNKAG QHDEPVHTAA VRIQNKFRSW KGRRDYLLIR QRIIKIQAHV 960 RGHQVRNKYK NIIWSVGILE KVILRWRRKG SGLRGFKPEA TLTEGSNMQD RPVQEDDYDF 1020 LKEGRKQTEQ RLQKALARVK SMVQYPEARD QYRRLLNVVS DMKDTTTTSD GAPSNSGEAA 1080 DFGDDLIDLD DLLDDDTFMS TAP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
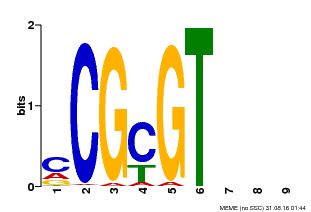
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF253511 | 0.0 | AF253511.1 Nicotiana tabacum anther ethylene-upregulated protein ER1 (ER1) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009763882.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 3-like isoform X1 | ||||
| Refseq | XP_016478076.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 3-like isoform X1 | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | A0A1S4AN37 | 0.0 | A0A1S4AN37_TOBAC; calmodulin-binding transcription activator 3-like isoform X1 | ||||
| TrEMBL | A0A1U7V6U6 | 0.0 | A0A1U7V6U6_NICSY; calmodulin-binding transcription activator 3-like isoform X1 | ||||
| STRING | XP_009763882.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2230 | 21 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



