 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_015894025.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rhamnaceae; Paliureae; Ziziphus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 513aa MW: 57065.8 Da PI: 6.262 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.9 | 1.9e-32 | 278 | 332 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
+pr+rWtpeLHerFveav++L G+ekAtPk +l+lm+v gLt++hvkSHLQkYRl
XP_015894025.1 278 RPRMRWTPELHERFVEAVNKLDGAEKATPKGVLKLMNVGGLTIYHVKSHLQKYRL 332
59****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 11.181 | 275 | 335 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.58E-18 | 276 | 332 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 5.5E-31 | 277 | 333 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.3E-24 | 279 | 333 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.5E-9 | 280 | 331 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 7.0E-23 | 392 | 439 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 513 aa Download sequence Send to blast |
MKVGLPSTTM NRHNVISITQ TEAVKGGTQP YCSALSPVHD LLNVQSESQR LFSSECSSAH 60 PSPFIRAESL SSPTHMQESS IQPQKYSFRS DSNSSVSPGS NVQHPKSEFS RSSVFCTSLY 120 QSSSTSSETS RQLGNLPFLP HPSTCNQPIS AAVSTKSTLL YSGDLSDQYD EEQSEALMKD 180 FLNLSADSSN GSFHHGFGCA SDSLALTEQL ELHFLSDELD IAITDHGENP GLDEIYETPQ 240 PSSKPALGLT CNQSSRATTP SVDELSNHLS SGTATAHRPR MRWTPELHER FVEAVNKLDG 300 AEKATPKGVL KLMNVGGLTI YHVKSHLQKY RLAKYMPEKK EGEEQMIERF YACLKKSETK 360 FILLLASPKC IDICIKYQCR FYEIHLSCHR SVQITEALRM QMEVQKQLHE QLEVQRALQL 420 RIEEHARYLQ KILEEQQKAG SALESTQALS SLSDSEHQPS PLAESAKSDS SSPKRLKQKA 480 TESNELECTK RLRLEDKPES ARDDIAAVEN PEQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 4e-25 | 278 | 335 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 4e-25 | 278 | 335 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 4e-25 | 278 | 335 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 4e-25 | 278 | 335 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 4e-25 | 278 | 335 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 4e-25 | 278 | 335 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 4e-25 | 278 | 335 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 4e-25 | 278 | 335 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00354 | DAP | Transfer from AT3G13040 | Download |
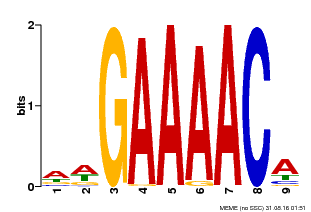
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_015894025.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JN714378 | 1e-138 | JN714378.1 Ziziphus mauritiana microsatellite Zam192 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024934181.1 | 0.0 | myb family transcription factor PHL6 | ||||
| TrEMBL | A0A2P5FLT0 | 0.0 | A0A2P5FLT0_TREOI; Octamer-binding transcription factor | ||||
| STRING | EMJ16523 | 0.0 | (Prunus persica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF13469 | 19 | 25 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G13040.2 | 1e-115 | G2-like family protein | ||||



