 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_015880907.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rhamnaceae; Paliureae; Ziziphus
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 846aa MW: 97432.8 Da PI: 6.8564 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 85.1 | 1.1e-26 | 89 | 191 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk............tekerrtraetrtgCkaklkvkkekdgkwevtklel 83
+fY+eYA+++GF++ +++s++sk+++e+++++f Cs++g+++e +k+ e++ +r+ ++t+Cka+++vk++ dgkw+++++++
XP_015880907.1 89 SFYQEYARSMGFNTAIQNSRRSKTSREFIDAKFACSRYGTKREYDKSfnrprsrqnkqdPENATGRRSCSKTDCKASMHVKRRPDGKWVIHNFVK 183
5*******************************************9999999988766554555559999************************** PP
FAR1 84 eHnHelap 91
eHnHel p
XP_015880907.1 184 EHNHELLP 191
*****975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 1.6E-24 | 89 | 191 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 3.7E-32 | 289 | 381 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.025 | 569 | 605 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 1.8E-4 | 579 | 602 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.0E-7 | 580 | 607 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 846 aa Download sequence Send to blast |
MDIDLRLPSG EHDKEGEEPT GIDNMLDNEE KLHNGDIETG NMVDIVDDVR AEDGGDLNSP 60 TTDIVVFKED TNLEPLSGME FESHGEAYSF YQEYARSMGF NTAIQNSRRS KTSREFIDAK 120 FACSRYGTKR EYDKSFNRPR SRQNKQDPEN ATGRRSCSKT DCKASMHVKR RPDGKWVIHN 180 FVKEHNHELL PAQAVSEQTR KMYAAMARQF AEYKNVVGLK NDLKSPFDKG RNLALEAGDL 240 KNLLDFCTQM QNMNSDFFYA IDLGEDQRLK NVFWVDAKSR HDYTNFNDVV SFDTTYIRNK 300 YKMPLALFVG VNQHYQFMLL GCALVSDESA TTFSWLMQTW LKAMGGQAPK VIITDHDKAI 360 KSVIPDIFPS AHHYFCLWHM MGKVTENLGH VIKRHENFIA KFEKCIHRSW TIEEFEKRWW 420 KILEKFELKE DEWMQLLYED RKQWVPTFMR DAFLAGMSTV QRSESVNCFF DKYVHKKTTV 480 QEFLKQYEAI LQDRYEEEAK ADSDTWNKQP TLKSPSPLEK SVSGVYTHAV FKKFQVEVLG 540 AVACHPKRER QDETGITFRV QDFERNLDFI VLWNEMKSEV SCLCRLFEYK GYLCRHAMIV 600 LQICGLSAIP SQYILKRWTK DAKNRHLTGE ESDHLQSRVQ RYNDLCQRAI KLTEEGSISQ 660 ESYSIACRAL DEAFSNCVSV NNSSKSLVEA STSTPHGLLC IEEDNQNKSM GKQNKKKNPT 720 KKRKVSFEPD VMAVGAQDSL QQMDKLNSRT VTLDGYYGAQ QSVQGMVQLN LMAPTRDNYY 780 GNQQTIQGLG QLNSIAPSHD GYYGTQQSMH GLGQMDFFRA PGFAYNIRDD PNVRTAALHD 840 DTSRHG |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
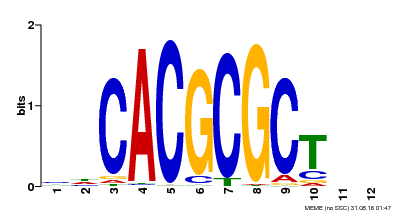
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_015880907.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015880907.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A251Q297 | 0.0 | A0A251Q297_PRUPE; Uncharacterized protein | ||||
| STRING | XP_008229656.1 | 0.0 | (Prunus mume) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF40 | 30 | 530 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||



