 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_015879435.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rhamnaceae; Paliureae; Ziziphus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 295aa MW: 33593.3 Da PI: 6.802 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.4 | 2.5e-18 | 86 | 138 | 4 | 56 |
-SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 4 RttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+ +++ eq+++Le+ Fe+ +++ e++ +LAk lgL+ rq+ +WFqNrRa++k
XP_015879435.1 86 KKRLNLEQVKTLEKSFEMGNKLEPERKMQLAKALGLQPRQIAIWFQNRRARWK 138
4568889*********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 129.6 | 1.2e-41 | 84 | 176 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
ekk+rl+ eqvk+LE+sFe+ +kLeperK++la++Lglqprq+a+WFqnrRAR+ktkqlEkdye+Lk+++dal+++n++L +++++L++el++
XP_015879435.1 84 EKKKRLNLEQVKTLEKSFEMGNKLEPERKMQLAKALGLQPRQIAIWFQNRRARWKTKQLEKDYEVLKKQFDALRSDNDALLAQNKKLHAELST 176
69**************************************************************************************99975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.88E-19 | 76 | 142 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.022 | 80 | 140 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.2E-17 | 83 | 144 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 5.96E-16 | 85 | 141 | No hit | No description |
| Pfam | PF00046 | 1.6E-15 | 86 | 138 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 3.0E-19 | 88 | 147 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 1.1E-5 | 111 | 120 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 115 | 138 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.1E-5 | 120 | 136 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 6.2E-16 | 140 | 180 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 295 aa Download sequence Send to blast |
MAFQPHSFMF QAHEDHQDLH LSSSASLNTL HFHAGGGHGA MPFMMKRSLS FSGVHNHHEN 60 IEQCEEVHGG GDDDLSDDGS QIGEKKKRLN LEQVKTLEKS FEMGNKLEPE RKMQLAKALG 120 LQPRQIAIWF QNRRARWKTK QLEKDYEVLK KQFDALRSDN DALLAQNKKL HAELSTLKGR 180 DSNEAGLSNL KKENEGSWSN GSENSSDINL EISRTPVINS PASSQINGKN FFPSTLRPSS 240 ITQFLQGTSR SDIQGLKVDQ HHHHQMVQEE SFCNMFNGIE DQQGFWPWPE QHQFH |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 132 | 140 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
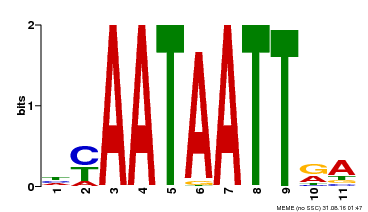
| Motif ID | Method | Source | Motif file |
| MP00322 | DAP | Transfer from AT3G01220 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_015879435.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. {ECO:0000269|PubMed:12644682}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015879435.1 | 0.0 | homeobox-leucine zipper protein ATHB-20-like | ||||
| Swissprot | Q8LAT0 | 9e-85 | ATB20_ARATH; Homeobox-leucine zipper protein ATHB-20 | ||||
| TrEMBL | A0A251Q638 | 1e-142 | A0A251Q638_PRUPE; Uncharacterized protein | ||||
| STRING | EMJ17483 | 1e-142 | (Prunus persica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1271 | 33 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G01220.1 | 2e-76 | homeobox protein 20 | ||||



