 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_013630261.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 317aa MW: 35290.9 Da PI: 4.8461 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 66.6 | 4.8e-21 | 104 | 154 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s+y+G+r+++ +g+W+AeIrdp+e r++lg+f taeeAa+a++aa+++++g
XP_013630261.1 104 SQYRGIRQRP-WGKWAAEIRDPRE---GSRVWLGTFKTAEEAARAYDAAARRIRG 154
78********.**********965...3*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 9.43E-18 | 104 | 163 | No hit | No description |
| SuperFamily | SSF54171 | 1.5E-22 | 104 | 163 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 1.5E-13 | 104 | 154 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 2.6E-33 | 105 | 163 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 24.408 | 105 | 162 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 6.4E-39 | 105 | 168 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 7.4E-12 | 106 | 117 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 7.4E-12 | 128 | 144 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0070483 | Biological Process | detection of hypoxia | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005886 | Cellular Component | plasma membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 317 aa Download sequence Send to blast |
MCGGAIISDF IPPPRSRRVT SEFLWPDLKK SAKKRSSFFD LDDEFEADFQ GFKDDSSIDC 60 DDAKPFVFAG ARKPAVSAAT ADSVFGKKVV DGEGERSAKR KRKSQYRGIR QRPWGKWAAE 120 IRDPREGSRV WLGTFKTAEE AARAYDAAAR RIRGSKAKVN FPEEKENPPA KKVAPNPSPV 180 LAQNLDNSFD NMCFMEEKHQ VNNNNNQFGG NGYHQYFSSD QGSNSFGCSE FGWNDQAPIT 240 PEISSAFINN NSVTFAEEAD PAKQLKVMDF ETTYNSTEWD SSLDFFSGDA VATQDNGANP 300 MELWSIDEIN SMIGGVF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 8e-22 | 105 | 162 | 5 | 63 | ATERF1 |
| 3gcc_A | 8e-22 | 105 | 162 | 5 | 63 | ATERF1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Bol.1241 | 0.0 | leaf | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed in seedlings, leaves, flowers, siliques and germinating seeds. {ECO:0000269|PubMed:22020282}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in the activation of hypoxic gene expression and in ethylene response. Partially redundant with RAP2-2. Acts as a downstream regulator in the ethylene signaling pathway. {ECO:0000269|PubMed:22020282, ECO:0000269|PubMed:22530619}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00204 | DAP | Transfer from AT1G53910 | Download |
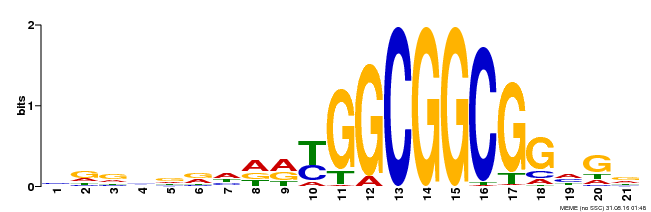
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_013630261.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by hypoxia but not by ethylene. {ECO:0000269|PubMed:22020282}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK353211 | 1e-70 | AK353211.1 Thellungiella halophila mRNA, complete cds, clone: RTFL01-22-M23. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013630261.1 | 0.0 | PREDICTED: ethylene-responsive transcription factor RAP2-12 | ||||
| Swissprot | Q9SSA8 | 1e-153 | RA212_ARATH; Ethylene-responsive transcription factor RAP2-12 | ||||
| TrEMBL | A0A0D3BMZ6 | 0.0 | A0A0D3BMZ6_BRAOL; Uncharacterized protein | ||||
| STRING | Bo3g184310.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM10 | 28 | 1650 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G53910.2 | 1e-137 | related to AP2 12 | ||||



